diff --git a/LLaVA-Med/.gitignore b/LLaVA-Med/.gitignore
deleted file mode 100644
index dcdbd68e77aae618467b9fd7c2d70591fa39ebef..0000000000000000000000000000000000000000
--- a/LLaVA-Med/.gitignore
+++ /dev/null
@@ -1,3 +0,0 @@
-__pycache__
-*.pyc
-*.egg-info
\ No newline at end of file
diff --git a/LLaVA-Med/CODE_OF_CONDUCT.md b/LLaVA-Med/CODE_OF_CONDUCT.md
deleted file mode 100644
index f9ba8cf65f3e3104dd061c178066ec8247811f33..0000000000000000000000000000000000000000
--- a/LLaVA-Med/CODE_OF_CONDUCT.md
+++ /dev/null
@@ -1,9 +0,0 @@
-# Microsoft Open Source Code of Conduct
-
-This project has adopted the [Microsoft Open Source Code of Conduct](https://opensource.microsoft.com/codeofconduct/).
-
-Resources:
-
-- [Microsoft Open Source Code of Conduct](https://opensource.microsoft.com/codeofconduct/)
-- [Microsoft Code of Conduct FAQ](https://opensource.microsoft.com/codeofconduct/faq/)
-- Contact [opencode@microsoft.com](mailto:opencode@microsoft.com) with questions or concerns
diff --git a/LLaVA-Med/LICENSE b/LLaVA-Med/LICENSE
deleted file mode 100644
index 60b9d4ee3f3e345ea4bbe33ece5b304a22a6257f..0000000000000000000000000000000000000000
--- a/LLaVA-Med/LICENSE
+++ /dev/null
@@ -1,62 +0,0 @@
-MICROSOFT RESEARCH LICENSE TERMS
-
-IF YOU LIVE IN THE UNITED STATES, PLEASE READ THE “BINDING ARBITRATION AND CLASS ACTION WAIVER” SECTION BELOW. IT AFFECTS HOW DISPUTES ARE RESOLVED.
-
-These license terms are an agreement between you and Microsoft Corporation (or one of its affiliates). They apply to the source code, object code, machine learning models, or data (collectively “Materials”) that accompany this license. IF YOU COMPLY WITH THESE LICENSE TERMS, YOU HAVE THE RIGHTS BELOW. BY USING THE MATERIALS, YOU ACCEPT THESE TERMS.
-
-1) INSTALLATION AND USE RIGHTS TO THE MATERIALS.
-
-Subject to the terms of this agreement, you have the below rights, if applicable, to use the Materials solely for non-commercial, non-revenue generating, research purposes:
-
- a) Source Code. If source code is included, you may use and modify the source code, but you may not distribute the source code.
- b) Object Code. If object code is included, you may use the object code, but you may not distribute the object code.
- c) Models. If machine learning model(s) are included, you may use the model(s), but you may not distribute the models.
- d) Data. If data is included, you may use and modify the data, but your use and modification must be consistent with the consent under which the data was provided and/or gathered and you may not distribute the data or your modifications to the data.
-
-2) SCOPE OF LICENSE. The Materials are licensed, not sold. Microsoft reserves all other rights. Unless applicable law gives you more rights despite this limitation, you will not (and have no right to):
-
- a) work around any technical limitations in the Materials that only allow you to use it in certain ways;
- b) reverse engineer, decompile or disassemble the Materials;
- c) remove, minimize, block, or modify any notices of Microsoft or its suppliers in the Materials;
- d) use the Materials in any way that is against the law or to create or propagate malware; or
- e) share, publish, distribute or lend the Materials, provide the Materials as a stand-alone hosted solution for others to use, or transfer the Materials or this agreement to any third party.
-
-3) PERSONAL DATA. If the data (set forth in Section 1(c) above) includes or is found to include any data that enables any ability to identify an individual (“Personal Data”), you will not use such Personal Data for any purpose other than was authorized and consented to by the data subject/research participant. You will not use Personal Data to contact any person. You will keep Personal Data in strict confidence. You will not share any Personal Data that is collected or in your possession with any third party for any reason and as required under the original consent agreement. Further, you will destroy the Personal Data and any backup or copies, immediately upon the completion of your research.
-
-4) LICENSE TO MICROSOFT. Notwithstanding the limitations in Section 1, you may distribute your modifications back to Microsoft, and if you do provide Microsoft with modifications of the Materials, you hereby grant Microsoft, without any restrictions or limitations, a non-exclusive, perpetual, irrevocable, royalty-free, assignable and sub-licensable license, to reproduce, publicly perform or display, install, use, modify, post, distribute, make and have made, sell and transfer such modifications and derivatives for any purpose.
-
-5) PUBLICATION. You may publish (or present papers or articles) on your results from using the Materials provided that no material or substantial portion of the Materials is included in any such publication or presentation.
-
-6) FEEDBACK. Any feedback about the Materials provided by you to us is voluntarily given, and Microsoft shall be free to use the feedback as it sees fit without obligation or restriction of any kind, even if the feedback is designated by you as confidential. Such feedback shall be considered a contribution and licensed to Microsoft under the terms of Section 4 above.
-
-7) COMPLIANCE WITH TRADE LAWS. You acknowledge that the Materials may be subject to applicable trade laws in one or more countries. You will comply with all relevant laws and regulations applicable to the import or export of the Materials, including but not limited to, trade laws such as the U.S. Export Administration Regulations or other end-user, end use, and destination restrictions by the U.S. and other governments, as well as sanctions regulations administered by the U.S. Office of Foreign Assets Control. Microsoft may suspend or terminate the agreement immediately to the extent that Microsoft reasonably concludes that continued performance would violate trade laws or put it at risk of becoming subject to sanctions or penalties under trade laws. For additional information, see www.microsoft.com/exporting.
-
-8) SUPPORT SERVICES. Microsoft is not obligated under this agreement to provide any support services for the Materials. Any support provided is “as is”, “with all faults”, and without warranty of any kind.
-
-9) BINDING ARBITRATION AND CLASS ACTION WAIVER. This Section applies if you live in (or, if a business, your principal place of business is in) the United States. If you and Microsoft have a dispute, you and Microsoft agree to try for 60 days to resolve it informally. If you and Microsoft can’t, you and Microsoft agree to binding individual arbitration before the American Arbitration Association under the Federal Arbitration Act (“FAA”), and not to sue in court in front of a judge or jury. Instead, a neutral arbitrator will decide. Class action lawsuits, class-wide arbitrations, private attorney-general actions, and any other proceeding where someone acts in a representative capacity are not allowed; nor is combining individual proceedings without the consent of all parties. The complete Arbitration Agreement contains more terms and is at aka.ms/arb-agreement-1. You and Microsoft agree to these terms.
-
-10) ENTIRE AGREEMENT. This agreement, and any other terms Microsoft may provide for supplements, updates, or third-party applications, is the entire agreement for the Materials.
-
-11) APPLICABLE LAW AND PLACE TO RESOLVE DISPUTES. If you acquired the Materials in the United States or Canada, the laws of the state or province where you live (or, if a business, where your principal place of business is located) govern the interpretation of this agreement, claims for its breach, and all other claims (including consumer protection, unfair competition, and tort claims), regardless of conflict of laws principles, except that the FAA governs everything related to arbitration. If you acquired the Materials in any other country, its laws apply, except that the FAA governs everything related to arbitration. If U.S. federal jurisdiction exists, you and Microsoft consent to exclusive jurisdiction and venue in the federal court in King County, Washington for all disputes heard in court (excluding arbitration). If not, you and Microsoft consent to exclusive jurisdiction and venue in the Superior Court of King County, Washington for all disputes heard in court (excluding arbitration).
-
-12) CONSUMER RIGHTS; REGIONAL VARIATIONS. This agreement describes certain legal rights. You may have other rights, including consumer rights, under the laws of your state, province, or country. Separate and apart from your relationship with Microsoft, you may also have rights with respect to the party from which you acquired the Materials. This agreement does not change those other rights if the laws of your state, province, or country do not permit it to do so. For example, if you acquired the Materials in one of the below regions, or mandatory country law applies, then the following provisions apply to you:
-
- a) Australia. You have statutory guarantees under the Australian Consumer Law and nothing in this agreement is intended to affect those rights.
-
- b) Canada. If you acquired this software in Canada, you may stop receiving updates by turning off the automatic update feature, disconnecting your device from the Internet (if and when you re-connect to the Internet, however, the Materials will resume checking for and installing updates), or uninstalling the Materials. The product documentation, if any, may also specify how to turn off updates for your specific device or software.
-
- c) Germany and Austria.
-
- i. Warranty. The properly licensed software will perform substantially as described in any Microsoft materials that accompany the Materials. However, Microsoft gives no contractual guarantee in relation to the licensed software.
-
- ii. Limitation of Liability. In case of intentional conduct, gross negligence, claims based on the Product Liability Act, as well as, in case of death or personal or physical injury, Microsoft is liable according to the statutory law.
-
- Subject to the foregoing clause (ii), Microsoft will only be liable for slight negligence if Microsoft is in breach of such material contractual obligations, the fulfillment of which facilitate the due performance of this agreement, the breach of which would endanger the purpose of this agreement and the compliance with which a party may constantly trust in (so-called "cardinal obligations"). In other cases of slight negligence, Microsoft will not be liable for slight negligence.
-
-13) DISCLAIMER OF WARRANTY. THE MATERIALS ARE LICENSED “AS IS.” YOU BEAR THE RISK OF USING THEM. MICROSOFT GIVES NO EXPRESS WARRANTIES, GUARANTEES, OR CONDITIONS. TO THE EXTENT PERMITTED UNDER APPLICABLE LAWS, MICROSOFT EXCLUDES ALL IMPLIED WARRANTIES, INCLUDING MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE, AND NON-INFRINGEMENT.
-
-14) LIMITATION ON AND EXCLUSION OF DAMAGES. IF YOU HAVE ANY BASIS FOR RECOVERING DAMAGES DESPITE THE PRECEDING DISCLAIMER OF WARRANTY, YOU CAN RECOVER FROM MICROSOFT AND ITS SUPPLIERS ONLY DIRECT DAMAGES UP TO U.S. $5.00. YOU CANNOT RECOVER ANY OTHER DAMAGES, INCLUDING CONSEQUENTIAL, LOST PROFITS, SPECIAL, INDIRECT OR INCIDENTAL DAMAGES.
-
-This limitation applies to (a) anything related to the Materials, services, content (including code) on third party Internet sites, or third party applications; and (b) claims for breach of contract, warranty, guarantee, or condition; strict liability, negligence, or other tort; or any other claim; in each case to the extent permitted by applicable law.
-
-It also applies even if Microsoft knew or should have known about the possibility of the damages. The above limitation or exclusion may not apply to you because your state, province, or country may not allow the exclusion or limitation of incidental, consequential, or other damages.
diff --git a/LLaVA-Med/README.md b/LLaVA-Med/README.md
deleted file mode 100644
index 5312c35bc11c1c8f44dafdddeee416ac59fcf739..0000000000000000000000000000000000000000
--- a/LLaVA-Med/README.md
+++ /dev/null
@@ -1,260 +0,0 @@
-# LLaVA-Med: Large Language and Vision Assistant for Biomedicine
-
-*Visual instruction tuning towards building large language and vision models with GPT-4 level capabilities in the biomedicine space.*
-
-[[Paper, NeurIPS 2023 Datasets and Benchmarks Track (Spotlight)](https://arxiv.org/abs/2306.00890)]
-
-**LLaVA-Med: Training a Large Language-and-Vision Assistant for Biomedicine in One Day**
-
-[Chunyuan Li*](https://chunyuan.li/), [Cliff Wong*](https://scholar.google.com/citations?user=Sl05ifcAAAAJ&hl=en), [Sheng Zhang*](https://scholar.google.com/citations?user=-LVEXQ8AAAAJ&hl=en), [Naoto Usuyama](https://www.microsoft.com/en-us/research/people/naotous/), [Haotian Liu](https://hliu.cc), [Jianwei Yang](https://jwyang.github.io/), [Tristan Naumann](https://scholar.google.com/citations?user=cjlSeqwAAAAJ&hl=en), [Hoifung Poon](https://scholar.google.com/citations?user=yqqmVbkAAAAJ&hl=en), [Jianfeng Gao](https://scholar.google.com/citations?user=CQ1cqKkAAAAJ&hl=en) (*Equal Contribution)
-
-
- 
-
- *Generated by GLIGEN using the grounded inpainting mode, with three boxes: ``white doctor coat``, ``stethoscope``, ``white doctor hat with a red cross sign``.*
-
-
-
-
-## Release
-
-- [May 13, 2024] 🔥LLaVA-Med v1.5 is out! It is not only significantly better (see the [evaluation results](docs/llava_med_performance.md#llava-med-15-performance).) but also much easier to use: no more *delta* weights! Now you can directly load our model from the [🤗 Hub](https://huggingface.co/microsoft/llava-med-v1.5-mistral-7b). The original LLaVA-Med (i.e., v1.0.0) codebase has been moved to [Archive](#archive).
-- [Nov 8, 2023] LLaVA-Med is open-sourced under the MSR release policy. Huge thanks to commitment of the team, and patience of the community.
-- [Sept, 2023] LLaVA-Med is accepted in NeurIPS 2023 Datasets and Benchmarks Track, as a spotlight presentation.
-- [June 1, 2023] 🔥 We released **LLaVA-Med: Large Language and Vision Assistant for Biomedicine**, a step towards building biomedical domain large language and vision models with GPT-4 level capabilities. Checkout the [paper](https://arxiv.org/abs/2306.00890)
-
-
- 
-
- *LLaVA-Med was initialized with the general-domain LLaVA and then continuously trained in a curriculum learning fashion (first biomedical concept alignment then full-blown instruction-tuning). We evaluated LLaVA-Med on standard visual conversation and question answering tasks.*
-
-
-[](Research%20License.docx)
-[](https://creativecommons.org/licenses/by-nc/4.0/deed.en)
-**Usage and License Notices**: The data, code, and model checkpoints are intended and licensed for research use only. They are also subject to additional restrictions dictated by the Terms of Use: LLaMA, Vicuna and GPT-4 respectively. The data is made available under CC BY NC 4.0. The data, code, and model checkpoints may be used for non-commercial purposes and any models trained using the dataset should be used only for research purposes. It is expressly prohibited for models trained on this data to be used in clinical care or for any clinical decision making purposes.
-
-## Contents
-
-- [Install](#install)
-- [Model Download](#model-download)
-- [Serving](#serving)
-- [Evaluation](#evaluation)
-- [Data Download](#data-download)
-- [Archive](#archive)
-- [Model Description](#model-description)
-
-## Install
-
-1. Clone this repository and navigate to LLaVA-Med folder
-```bash
-https://github.com/microsoft/LLaVA-Med.git
-cd LLaVA-Med
-```
-
-2. Install Package: Create conda environment
-
-```Shell
-conda create -n llava-med python=3.10 -y
-conda activate llava-med
-pip install --upgrade pip # enable PEP 660 support
-pip install -e .
-```
-
-## Model Download
-
-
- Model Descriptions | 🤗 Huggingface Hub |
-| --- | ---: |
-| LLaVA-Med v1.5 | [microsoft/llava-med-v1.5-mistral-7b](https://huggingface.co/microsoft/llava-med-v1.5-mistral-7b) |
-
-
-
-## Serving
-
-### Web UI
-
-#### Launch a controller
-```Shell
-python -m llava.serve.controller --host 0.0.0.0 --port 10000
-```
-
-#### Launch a model worker
-```Shell
-python -m llava.serve.model_worker --host 0.0.0.0 --controller http://localhost:10000 --port 40000 --worker http://localhost:40000 --model-path microsoft/llava-med-v1.5-mistral-7b --multi-modal
-```
-Wait until the process finishes loading the model and you see "Uvicorn running on ...".
-
-#### Launch a model worker (Multiple GPUs, when GPU VRAM <= 24GB)
-
-If your the VRAM of your GPU is less than 24GB (e.g., RTX 3090, RTX 4090, etc.), you may try running it with multiple GPUs.
-
-```Shell
-python -m llava.serve.model_worker --host 0.0.0.0 --controller http://localhost:10000 --port 40000 --worker http://localhost:40000 --model-path microsoft/llava-med-v1.5-mistral-7b --multi-modal --num-gpus 2
-```
-Wait until the process finishes loading the model and you see "Uvicorn running on ...".
-
-
-#### Send a test message
-```Shell
-python -m llava.serve.test_message --model-name llava-med-v1.5-mistral-7b --controller http://localhost:10000
-```
-
-#### Launch a gradio web server.
-```Shell
-python -m llava.serve.gradio_web_server --controller http://localhost:10000
-```
-#### You can open your browser and chat with a model now.
-
-
-## Evaluation
-
-### Medical Visual Chat (GPT-assisted Evaluation)
-
-Our GPT-assisted evaluation pipeline for multimodal modeling is provided for a comprehensive understanding of the capabilities of vision-language models. Please see our paper for more details.
-
-#### 1. Azure OpenAI Connection Info.
-
-Open [llava/eval/llm.py](llava/eval/llm.py?plain=1#L33) and insert your Azure OpenAI Endpoint and API KEY
-```Shell
-openai_cxn_dict = {
- 'default': {
- 'endpoint': "INSERT YOUR AZURE OPENAI ENDPOINT HERE",
- 'api_key': "INSERT YOUR AZURE OPENAI API KEY HERE",
- },
- }
-```
-* GPT-4 inference was only tested using Azure OpenAI API. If you are using OpenAI API, you need to replace [llava/eval/llm.py (line 55)](llava/eval/llm.py?plain=1#L55) AsyncAzureOpenAI with AsyncOpenAI.
-
-#### 2. Deployment ID
-In [llava/eval/eval_multimodal_chat_gpt_score.py (line 55)](llava/eval/eval_multimodal_chat_gpt_score.py?plain=1#L55), replace with your GPT-4 model deployment id if necessary:
-
-#### 3. Download Images
-
-```Shell
-wget https://hanoverprod.z21.web.core.windows.net/med_llava/multimodal_chat_eval/llava_med_test_image_urls.jsonl -P data/
-python llava/data/download_images.py \
- --input_path data/llava_med_test_image_urls.jsonl \
- --pmc_output_path data/pmc \
- --images_output_path data/images
-```
-
-#### 4. Multimodal Chat Inference
-In our case, [`llava_med_eval_qa50_qa.jsonl`](/data/eval/llava_med_eval_qa50_qa.jsonl) contains the questions, context (captions and inline-mentions) and responses generated by text-only GPT-4 (0314), which we treat as ground truth.
-
-```Shell
-PYTHONPATH=. python llava/eval/model_vqa.py \
- --conv-mode mistral_instruct \
- --model-path microsoft/llava-med-v1.5-mistral-7b \
- --question-file data/eval/llava_med_eval_qa50_qa.jsonl \
- --image-folder data/images \
- --answers-file /path/to/answer-file.jsonl \
- --temperature 0.0
-```
-
-#### 5. GPT-4 Evaluation of the Generated Answers
-
-```Shell
-python llava/eval/eval_multimodal_chat_gpt_score.py \
- --answers-file /path/to/answer-file.jsonl \
- --question-file data/eval/llava_med_eval_qa50_qa.jsonl \
- --scores-file /path/to/scores-file.jsonl
-```
-
-#### 6. Summarize the Evaluation Results
-
-```Shell
-python llava/eval/summarize_gpt_review.py \
- --scores-file /path/to/scores-file.jsonl
-```
-
-## Data Download
-
-### LLaVA-Med Dataset
-
-
- 
-
- *The data statistics of biomedical multimodal instruction-following data: (a,b) The root verb-noun pairs of instruction and responses, where the inner circle of the plot represents the root verb of the output response, and the outer circle represents the direct nouns. (c) The distribution of images and QA pairs on the five domains, one image is shown per domain.*
-
-
-### Data Download
-| Alignment data files | Size |
-| --- | ---: |
-| [llava_med_alignment_500k.json](https://hanoverprod.z21.web.core.windows.net/med_llava/alignment/llava_med_alignment_500k.json) | 341.52 MiB |
-
-| Instruction-Tuning data files | Size |
-| --- | ---: |
-| [llava_med_instruct_10k.json](https://hanoverprod.z21.web.core.windows.net/med_llava/instruct/llava_med_instruct_10k.json) | 19.24 MiB |
-| [llava_med_instruct_60k.json](https://hanoverprod.z21.web.core.windows.net/med_llava/instruct/llava_med_instruct_60k.json) | 84.65 MiB |
-| [llava_med_instruct_60k_inline_mention.json](https://hanoverprod.z21.web.core.windows.net/med_llava/instruct/llava_med_instruct_60k_inline_mention.json) | 83.61 MiB |
-| [llava_med_instruct_fig_captions.json](https://hanoverprod.z21.web.core.windows.net/med_llava/instruct/llava_med_instruct_fig_captions.json) | 161.39 MiB |
-
-| Evaluation files | Size |
-| --- | ---: |
-| [llava_med_eval_qa50_qa.jsonl](https://hanoverprod.z21.web.core.windows.net/med_llava/eval/llava_med_eval_qa50_qa.jsonl) | 256.18 KiB |
-| [llava_med_eval_qa50_fig_captions.json](https://hanoverprod.z21.web.core.windows.net/med_llava/eval/llava_med_eval_qa50_fig_captions.json) | 51.82 KiB |
-| [llava_med_qa50_instruct_caption_in_text_cleaned-60k-3epoch.json](https://hanoverprod.z21.web.core.windows.net/med_llava/eval/llava_med_qa50_instruct_caption_in_text_cleaned-60k-3epoch.json) | 100.97 KiB |
-
-| Image URLS | Size |
-| --- | ---: |
-| [llava_med_image_urls.jsonl](https://hanoverprod.z21.web.core.windows.net/med_llava/llava_med_image_urls.jsonl) | 122.82 MiB |
-
-[download_images.py](https://github.com/microsoft/LLaVA-Med/blob/v1.0.0/llava/data/download_images.py) is used to download the PMC articles using the above image_urls file and extract the images
-
-To download our langauge-image multimodal instruction-folllowing dataset, please run the following script:
-```bash
-sh download_data.sh
-```
-
-
-## Archive
-
-- [LLaVA-Med v1.0](https://github.com/microsoft/LLaVA-Med/tree/v1.0.0)
-
-## Model Description
-
-Large Language and Vision Assistant for bioMedicine (i.e., “LLaVA-Med”) is a large language and vision model trained using a curriculum learning method for adapting LLaVA to the biomedical domain. It is an open-source release intended for research use only to facilitate reproducibility of the corresponding paper which claims improved performance for open-ended biomedical questions answering tasks, including common visual question answering (VQA) benchmark datasets such as PathVQA and VQA-RAD.
-
-### Model Uses
-
-#### Intended Use
-
-The data, code, and model checkpoints are intended to be used solely for (I) future research on visual-language processing and (II) reproducibility of the experimental results reported in the reference paper. The data, code, and model checkpoints are not intended to be used in clinical care or for any clinical decision making purposes.
-
-#### Primary Intended Use
-
-The primary intended use is to support AI researchers reproducing and building on top of this work. LLaVA-Med and its associated models should be helpful for exploring various biomedical vision-language processing (VLP ) and vision question answering (VQA) research questions.
-
-#### Out-of-Scope Use
-
-**Any** deployed use case of the model --- commercial or otherwise --- is out of scope. Although we evaluated the models using a broad set of publicly-available research benchmarks, the models and evaluations are intended *for research use only* and not intended for deployed use cases. Please refer to [the associated paper](https://aka.ms/llava-med) for more details.
-
-### Data
-
-This model builds upon [PMC-15M dataset](https://aka.ms/biomedclip-paper), which is a large-scale parallel image-text dataset for biomedical vision-language processing. It contains 15 million figure-caption pairs extracted from biomedical research articles in PubMed Central. It covers a diverse range of biomedical image types, such as microscopy, radiography, histology, and more.
-
-### Limitations
-
-This model was developed using English corpora, and thus may be considered English-only. This model is evaluated on a narrow set of biomedical benchmark tasks, described in [LLaVA-Med paper](https://aka.ms/llava-med). As such, it is not suitable for use in any clinical setting. Under some conditions, the model may make inaccurate predictions and display limitations, which may require additional mitigation strategies. In particular, this model is likely to carry many of the limitations of the model from which it is derived, [LLaVA](https://llava-vl.github.io/).
-
-Further, this model was developed in part using the [PMC-15M](https://aka.ms/biomedclip-paper) dataset. The figure-caption pairs that make up this dataset may contain biases reflecting the current practice of academic publication. For example, the corresponding papers may be enriched for positive findings, contain examples of extreme cases, and otherwise reflect distributions that are not representative of other sources of biomedical data.
-
-## Acknowledgement
-
-If you find LLaVA-Med useful for your your research and applications, please cite using this BibTeX:
-
-```bibtex
-@article{li2023llavamed,
- title={Llava-med: Training a large language-and-vision assistant for biomedicine in one day},
- author={Li, Chunyuan and Wong, Cliff and Zhang, Sheng and Usuyama, Naoto and Liu, Haotian and Yang, Jianwei and Naumann, Tristan and Poon, Hoifung and Gao, Jianfeng},
- journal={arXiv preprint arXiv:2306.00890},
- year={2023}
-}
-```
-
-
-## Related Projects
-
-- [LLaVA](https://llava-vl.github.io/)
-- [BiomedCLIP](https://huggingface.co/microsoft/BiomedCLIP-PubMedBERT_256-vit_base_patch16_224)
-- [Instruction Tuning with GPT-4](https://github.com/Instruction-Tuning-with-GPT-4/GPT-4-LLM)
diff --git a/LLaVA-Med/SECURITY.md b/LLaVA-Med/SECURITY.md
deleted file mode 100644
index f7b89984f0fb5dd204028bc525e19eefc0859f4f..0000000000000000000000000000000000000000
--- a/LLaVA-Med/SECURITY.md
+++ /dev/null
@@ -1,41 +0,0 @@
-
-
-## Security
-
-Microsoft takes the security of our software products and services seriously, which includes all source code repositories managed through our GitHub organizations, which include [Microsoft](https://github.com/Microsoft), [Azure](https://github.com/Azure), [DotNet](https://github.com/dotnet), [AspNet](https://github.com/aspnet), [Xamarin](https://github.com/xamarin), and [our GitHub organizations](https://opensource.microsoft.com/).
-
-If you believe you have found a security vulnerability in any Microsoft-owned repository that meets [Microsoft's definition of a security vulnerability](https://docs.microsoft.com/en-us/previous-versions/tn-archive/cc751383(v=technet.10)), please report it to us as described below.
-
-## Reporting Security Issues
-
-**Please do not report security vulnerabilities through public GitHub issues.**
-
-Instead, please report them to the Microsoft Security Response Center (MSRC) at [https://msrc.microsoft.com/create-report](https://msrc.microsoft.com/create-report).
-
-If you prefer to submit without logging in, send email to [secure@microsoft.com](mailto:secure@microsoft.com). If possible, encrypt your message with our PGP key; please download it from the [Microsoft Security Response Center PGP Key page](https://www.microsoft.com/en-us/msrc/pgp-key-msrc).
-
-You should receive a response within 24 hours. If for some reason you do not, please follow up via email to ensure we received your original message. Additional information can be found at [microsoft.com/msrc](https://www.microsoft.com/msrc).
-
-Please include the requested information listed below (as much as you can provide) to help us better understand the nature and scope of the possible issue:
-
- * Type of issue (e.g. buffer overflow, SQL injection, cross-site scripting, etc.)
- * Full paths of source file(s) related to the manifestation of the issue
- * The location of the affected source code (tag/branch/commit or direct URL)
- * Any special configuration required to reproduce the issue
- * Step-by-step instructions to reproduce the issue
- * Proof-of-concept or exploit code (if possible)
- * Impact of the issue, including how an attacker might exploit the issue
-
-This information will help us triage your report more quickly.
-
-If you are reporting for a bug bounty, more complete reports can contribute to a higher bounty award. Please visit our [Microsoft Bug Bounty Program](https://microsoft.com/msrc/bounty) page for more details about our active programs.
-
-## Preferred Languages
-
-We prefer all communications to be in English.
-
-## Policy
-
-Microsoft follows the principle of [Coordinated Vulnerability Disclosure](https://www.microsoft.com/en-us/msrc/cvd).
-
-
\ No newline at end of file
diff --git a/LLaVA-Med/SUPPORT.md b/LLaVA-Med/SUPPORT.md
deleted file mode 100644
index dc72f0e5a0bc2807bf3df31dbc7455e6991b127a..0000000000000000000000000000000000000000
--- a/LLaVA-Med/SUPPORT.md
+++ /dev/null
@@ -1,25 +0,0 @@
-# TODO: The maintainer of this repo has not yet edited this file
-
-**REPO OWNER**: Do you want Customer Service & Support (CSS) support for this product/project?
-
-- **No CSS support:** Fill out this template with information about how to file issues and get help.
-- **Yes CSS support:** Fill out an intake form at [aka.ms/spot](https://aka.ms/spot). CSS will work with/help you to determine next steps. More details also available at [aka.ms/onboardsupport](https://aka.ms/onboardsupport).
-- **Not sure?** Fill out a SPOT intake as though the answer were "Yes". CSS will help you decide.
-
-*Then remove this first heading from this SUPPORT.MD file before publishing your repo.*
-
-# Support
-
-## How to file issues and get help
-
-This project uses GitHub Issues to track bugs and feature requests. Please search the existing
-issues before filing new issues to avoid duplicates. For new issues, file your bug or
-feature request as a new Issue.
-
-For help and questions about using this project, please **REPO MAINTAINER: INSERT INSTRUCTIONS HERE
-FOR HOW TO ENGAGE REPO OWNERS OR COMMUNITY FOR HELP. COULD BE A STACK OVERFLOW TAG OR OTHER
-CHANNEL. WHERE WILL YOU HELP PEOPLE?**.
-
-## Microsoft Support Policy
-
-Support for this **PROJECT or PRODUCT** is limited to the resources listed above.
diff --git a/LLaVA-Med/bitsandbytes-0.45.0-py3-none-manylinux_2_24_x86_64.whl b/LLaVA-Med/bitsandbytes-0.45.0-py3-none-manylinux_2_24_x86_64.whl
deleted file mode 100644
index 7de2e9a917992e764649667e2f295ad237d8718a..0000000000000000000000000000000000000000
--- a/LLaVA-Med/bitsandbytes-0.45.0-py3-none-manylinux_2_24_x86_64.whl
+++ /dev/null
@@ -1,3 +0,0 @@
-version https://git-lfs.github.com/spec/v1
-oid sha256:0f0323de1ff1fdf8383e79bdad1283516a4c05a6fd2b44a363bf4e059422305b
-size 69084267
diff --git a/LLaVA-Med/data/eval/llava_med_eval_qa50_qa.jsonl b/LLaVA-Med/data/eval/llava_med_eval_qa50_qa.jsonl
deleted file mode 100644
index 5896134a8534f954514e8f9760910d6e17bfb458..0000000000000000000000000000000000000000
--- a/LLaVA-Med/data/eval/llava_med_eval_qa50_qa.jsonl
+++ /dev/null
@@ -1,193 +0,0 @@
-{"question_id": 0, "image": "18063892_F3.jpg", "pair_id": "18063892_F3", "text": "What is the organ in the image?\n", "gpt4_answer": "The organ in the image is the conjunctiva, which is a thin, transparent tissue that covers the front part of the eye and lines the inside of the eyelids.", "fig_caption": "The gross histopathologic finding disclosed the portion of conjunctiva showing a cystic wall lined by a single layer of low cuboidal epithelium with edema and congestion measuring 0.1-0.2 ml in volume. (H & E stain, \u00d7100)", "in_text_mention": [{"tokens": "Inflammatory cells were not found in the vicinity ( Fig. 3 , 4 , 5 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 9, "end_tok": 11}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": true}, "type": "conversation", "fig_label": "Fig. 3"}
-{"question_id": 1, "image": "18063892_F3.jpg", "pair_id": "18063892_F3", "text": "What type of cells are lining the cystic wall?\n", "gpt4_answer": "The cystic wall is lined by a single layer of low cuboidal epithelium cells.", "fig_caption": "The gross histopathologic finding disclosed the portion of conjunctiva showing a cystic wall lined by a single layer of low cuboidal epithelium with edema and congestion measuring 0.1-0.2 ml in volume. (H & E stain, \u00d7100)", "in_text_mention": [{"tokens": "Inflammatory cells were not found in the vicinity ( Fig. 3 , 4 , 5 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 9, "end_tok": 11}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": true}, "type": "conversation", "fig_label": "Fig. 3"}
-{"question_id": 2, "image": "18063892_F3.jpg", "pair_id": "18063892_F3", "text": "Are there any inflammatory cells present?\n", "gpt4_answer": "No, there are no inflammatory cells found in the vicinity of the conjunctiva in this image.", "fig_caption": "The gross histopathologic finding disclosed the portion of conjunctiva showing a cystic wall lined by a single layer of low cuboidal epithelium with edema and congestion measuring 0.1-0.2 ml in volume. (H & E stain, \u00d7100)", "in_text_mention": [{"tokens": "Inflammatory cells were not found in the vicinity ( Fig. 3 , 4 , 5 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 9, "end_tok": 11}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": true}, "type": "conversation", "fig_label": "Fig. 3"}
-{"question_id": 3, "image": "18318598_pmed-0050054-g001.jpg", "pair_id": "18318598_pmed-0050054-g001", "text": "What is the difference between the Winnie and wild-type mice?\n", "gpt4_answer": "The histological differences between the Winnie and wild-type mice in the intestinal sections are related to the goblet cells and their mucin storage. In the Winnie mice, there are fewer goblet cells with smaller thecae (stored mucin) compared to the wild-type mice. Additionally, the Winnie goblet cells show the presence of PAS-positive/Alcian blue-negative accumulations, which are not observed in the wild-type mice. These differences suggest alterations in the mucin production and storage in the Winnie mice compared to the wild-type mice.", "fig_caption": "Histological Phenotype of Mice with Muc2 Mutations PAS/Alcian blue stained intestinal sections from Winnie and wild-type C57BL/6 mice. Note the reduced size of Alcian blue staining thecae (stored mucin) and the presence of PAS-positive/Alcian blue negative accumulations (arrows) in Winnie goblet cells. L, lumen.", "in_text_mention": [{"tokens": "Compared with wild-type littermates , Winnie small and large intestines were characterized by fewer goblet cells with smaller thecae , the presence of PAS positive Alcian blue negative material in the cytoplasm , and a reduction in secreted mucus ( Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 40, "end_tok": 42}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 4, "image": "18318598_pmed-0050054-g001.jpg", "pair_id": "18318598_pmed-0050054-g001", "text": "What is the purpose of the staining?\n", "gpt4_answer": "The purpose of the PAS (Periodic Acid-Schiff) and Alcian blue staining in this context is to visualize and differentiate specific components within the intestinal sections of the mice. PAS staining is used to detect polysaccharides, glycoproteins, and glycolipids, while Alcian blue staining is used to identify acidic mucopolysaccharides and mucins. By using these stains, researchers can better understand the differences in the goblet cells and mucin storage between the Winnie and wild-type mice, as well as observe any alterations in the intestinal tissue.", "fig_caption": "Histological Phenotype of Mice with Muc2 Mutations PAS/Alcian blue stained intestinal sections from Winnie and wild-type C57BL/6 mice. Note the reduced size of Alcian blue staining thecae (stored mucin) and the presence of PAS-positive/Alcian blue negative accumulations (arrows) in Winnie goblet cells. L, lumen.", "in_text_mention": [{"tokens": "Compared with wild-type littermates , Winnie small and large intestines were characterized by fewer goblet cells with smaller thecae , the presence of PAS positive Alcian blue negative material in the cytoplasm , and a reduction in secreted mucus ( Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 40, "end_tok": 42}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 5, "image": "18318598_pmed-0050054-g001.jpg", "pair_id": "18318598_pmed-0050054-g001", "text": "What is the role of goblet cells?\n", "gpt4_answer": "Goblet cells are specialized epithelial cells found in the lining of the respiratory, digestive, and reproductive tracts. Their primary function is to produce and secrete mucus, which is a viscous fluid composed of water, proteins, and glycoproteins called mucins. Mucus serves several important functions, such as lubricating and protecting the lining of the organs, trapping and removing foreign particles and pathogens, and maintaining the moisture and pH balance within the organ systems. Goblet cells play a crucial role in maintaining the health and integrity of the mucosal surfaces in the body.", "fig_caption": "Histological Phenotype of Mice with Muc2 Mutations PAS/Alcian blue stained intestinal sections from Winnie and wild-type C57BL/6 mice. Note the reduced size of Alcian blue staining thecae (stored mucin) and the presence of PAS-positive/Alcian blue negative accumulations (arrows) in Winnie goblet cells. L, lumen.", "in_text_mention": [{"tokens": "Compared with wild-type littermates , Winnie small and large intestines were characterized by fewer goblet cells with smaller thecae , the presence of PAS positive Alcian blue negative material in the cytoplasm , and a reduction in secreted mucus ( Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 40, "end_tok": 42}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 6, "image": "18652673_F3.jpg", "pair_id": "18652673_F3", "text": "What is the difference between the images of the depressed patient and the healthy control subject?\n", "gpt4_answer": "The difference between the images of the depressed patient and the healthy control subject is the presence of hypofrontality in the depressed patient. Hypofrontality refers to decreased activity in the prefrontal cortex (PFC), which can be observed as reduced cerebral blood flow (CBF) and glucose metabolism in the PFC, particularly in the dorsolateral prefrontal cortex (DLPFC). This is a characteristic finding in neuroimaging studies of depression.", "fig_caption": "Single photon emission computed tomography (SPECT) images from a depressed patient showing characteristic hypofrontality relative to a healthy control subject[82].", "in_text_mention": [{"tokens": "One of the most consistent findings in the neuroimaging of depression is decreased cerebral blood flow ( CBF ) and glucose metabolism in the PFC , particularly the DLPFC [ 77-85 ] ( figure 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 33, "end_tok": 35}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 7, "image": "18652673_F3.jpg", "pair_id": "18652673_F3", "text": "What imaging technique was used?\n", "gpt4_answer": "The imaging technique used in this case is single photon emission computed tomography (SPECT).", "fig_caption": "Single photon emission computed tomography (SPECT) images from a depressed patient showing characteristic hypofrontality relative to a healthy control subject[82].", "in_text_mention": [{"tokens": "One of the most consistent findings in the neuroimaging of depression is decreased cerebral blood flow ( CBF ) and glucose metabolism in the PFC , particularly the DLPFC [ 77-85 ] ( figure 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 33, "end_tok": 35}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 8, "image": "18652673_F3.jpg", "pair_id": "18652673_F3", "text": "What does hypofrontality indicate?\n", "gpt4_answer": "Hypofrontality indicates decreased activity in the prefrontal cortex (PFC) of the brain. In the context of depression, hypofrontality is associated with reduced cerebral blood flow (CBF) and glucose metabolism in the PFC, particularly in the dorsolateral prefrontal cortex (DLPFC). This finding suggests that there may be functional abnormalities in the PFC, which could be related to the cognitive and emotional symptoms experienced by individuals with depression. However, it is important to note that hypofrontality is not specific to depression and can be observed in other psychiatric and neurological conditions as well.", "fig_caption": "Single photon emission computed tomography (SPECT) images from a depressed patient showing characteristic hypofrontality relative to a healthy control subject[82].", "in_text_mention": [{"tokens": "One of the most consistent findings in the neuroimaging of depression is decreased cerebral blood flow ( CBF ) and glucose metabolism in the PFC , particularly the DLPFC [ 77-85 ] ( figure 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 33, "end_tok": 35}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 9, "image": "18673566_F1.jpg", "pair_id": "18673566_F1", "text": "What is the yellow waveform representing?\n", "gpt4_answer": "The yellow waveform represents the unfiltered, source-space projected, phase-locked average waveform resulting from a single equivalent current dipole. This waveform shows the M50, M100, and M200 responses, which are components of the brain's response to a stimulus.", "fig_caption": "Source space projection and time-frequency analysis. Left hemisphere data from a single participant are illustrated. A single equivalent current dipole was fit to the sensor data and is shown overlaid onto the co-registered MRI scan for the same individual (left). The yellow waveform is the unfiltered, source-space projected, phase-locked average waveform resulting from that dipole (the M50, M100 and M200 responses can be seen in the waveform). In the upper right panel, a time-frequency plot illustrates the transient gamma-band response in terms of power relative to the pre-stimulus baseline. The lower right panel illustrates the PLF for the same data. Note the peak power and PLF centered around 50 ms post-stimulus at 40 Hz.", "in_text_mention": [{"tokens": "A single equivalent current dipole ( ECD ) was then fit separately for left and right hemispheres using a spatiotemporal model to the observed data in the post-stimulus window between 40-80 ms , yielding parameter estimates of the x , y , and z ECD position information , as well as dipole orientation and magnitude over time ( see Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 59, "end_tok": 61}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 10, "image": "18673566_F1.jpg", "pair_id": "18673566_F1", "text": "What does the time-frequency plot illustrate?\n", "gpt4_answer": "The time-frequency plot illustrates the transient gamma-band response in terms of power relative to the pre-stimulus baseline. It shows how the power of the gamma-band activity changes over time in response to a stimulus.", "fig_caption": "Source space projection and time-frequency analysis. Left hemisphere data from a single participant are illustrated. A single equivalent current dipole was fit to the sensor data and is shown overlaid onto the co-registered MRI scan for the same individual (left). The yellow waveform is the unfiltered, source-space projected, phase-locked average waveform resulting from that dipole (the M50, M100 and M200 responses can be seen in the waveform). In the upper right panel, a time-frequency plot illustrates the transient gamma-band response in terms of power relative to the pre-stimulus baseline. The lower right panel illustrates the PLF for the same data. Note the peak power and PLF centered around 50 ms post-stimulus at 40 Hz.", "in_text_mention": [{"tokens": "A single equivalent current dipole ( ECD ) was then fit separately for left and right hemispheres using a spatiotemporal model to the observed data in the post-stimulus window between 40-80 ms , yielding parameter estimates of the x , y , and z ECD position information , as well as dipole orientation and magnitude over time ( see Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 59, "end_tok": 61}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 11, "image": "18673566_F1.jpg", "pair_id": "18673566_F1", "text": "What does the lower right panel show?\n", "gpt4_answer": "The lower right panel illustrates the Phase Locking Factor (PLF) for the same data. PLF is a measure of the consistency of the phase of a specific frequency component across trials. In this case, it shows the consistency of the phase of the gamma-band activity in response to the stimulus. The peak power and PLF are centered around 50 ms post-stimulus at 40 Hz.", "fig_caption": "Source space projection and time-frequency analysis. Left hemisphere data from a single participant are illustrated. A single equivalent current dipole was fit to the sensor data and is shown overlaid onto the co-registered MRI scan for the same individual (left). The yellow waveform is the unfiltered, source-space projected, phase-locked average waveform resulting from that dipole (the M50, M100 and M200 responses can be seen in the waveform). In the upper right panel, a time-frequency plot illustrates the transient gamma-band response in terms of power relative to the pre-stimulus baseline. The lower right panel illustrates the PLF for the same data. Note the peak power and PLF centered around 50 ms post-stimulus at 40 Hz.", "in_text_mention": [{"tokens": "A single equivalent current dipole ( ECD ) was then fit separately for left and right hemispheres using a spatiotemporal model to the observed data in the post-stimulus window between 40-80 ms , yielding parameter estimates of the x , y , and z ECD position information , as well as dipole orientation and magnitude over time ( see Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 59, "end_tok": 61}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 12, "image": "19838397_F0003.jpg", "pair_id": "19838397_F0003", "text": "What does the chest X-ray show?\n", "gpt4_answer": "The chest X-ray appears to be normal, with no signs of metastatic lesions or any other abnormalities.", "fig_caption": "Chest X-ray: Normal with no signs of metastatic lesions", "in_text_mention": [{"tokens": "Chest radiograph [ Figure 3 ] , ultrasound scan and computed tomography ( CT ) chest , abdomen , and pelvis [ Figure 4 ] did not reveal any metastases or lymph node involvement .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 3, "end_tok": 5}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 13, "image": "19838397_F0003.jpg", "pair_id": "19838397_F0003", "text": "Are there any signs of lymph node involvement?\n", "gpt4_answer": "Based on the chest X-ray, there are no signs of lymph node involvement.", "fig_caption": "Chest X-ray: Normal with no signs of metastatic lesions", "in_text_mention": [{"tokens": "Chest radiograph [ Figure 3 ] , ultrasound scan and computed tomography ( CT ) chest , abdomen , and pelvis [ Figure 4 ] did not reveal any metastases or lymph node involvement .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 3, "end_tok": 5}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 14, "image": "19838397_F0003.jpg", "pair_id": "19838397_F0003", "text": "What about metastases?\n", "gpt4_answer": "The chest X-ray does not show any signs of metastases. However, it is important to note that other imaging techniques, such as ultrasound scans and computed tomography (CT) scans, may provide more detailed information about the presence or absence of metastases or lymph node involvement in other areas of the body.", "fig_caption": "Chest X-ray: Normal with no signs of metastatic lesions", "in_text_mention": [{"tokens": "Chest radiograph [ Figure 3 ] , ultrasound scan and computed tomography ( CT ) chest , abdomen , and pelvis [ Figure 4 ] did not reveal any metastases or lymph node involvement .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 3, "end_tok": 5}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 15, "image": "20406459_F5.jpg", "pair_id": "20406459_F5", "text": "What organ is shown in the image?\n", "gpt4_answer": "The image shows a gross specimen of a left renal mass, which is related to the kidney.", "fig_caption": "Left renal mass gross specimen demonstrating the renal venous thrombus.", "in_text_mention": [{"tokens": "Final pathology for case 2 was primary tumor size 7.6 cm , RCC , clear cell type , Fuhrman grade 3 , stage T3bNxM1 , with negative margins ( Figure 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 29, "end_tok": 31}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 16, "image": "20406459_F5.jpg", "pair_id": "20406459_F5", "text": "What is the main feature in the image?\n", "gpt4_answer": "The main feature in the image is the renal venous thrombus, which is a blood clot that has formed within the renal vein.", "fig_caption": "Left renal mass gross specimen demonstrating the renal venous thrombus.", "in_text_mention": [{"tokens": "Final pathology for case 2 was primary tumor size 7.6 cm , RCC , clear cell type , Fuhrman grade 3 , stage T3bNxM1 , with negative margins ( Figure 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 29, "end_tok": 31}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 17, "image": "20406459_F5.jpg", "pair_id": "20406459_F5", "text": "What type of tumor is mentioned in the context?\n", "gpt4_answer": "The tumor mentioned in the context is a renal cell carcinoma (RCC), which is a type of kidney cancer. It is specifically the clear cell type and has a Fuhrman grade of 3. The stage of the tumor is T3bNxM1, indicating that it has advanced and may have spread to other parts of the body.", "fig_caption": "Left renal mass gross specimen demonstrating the renal venous thrombus.", "in_text_mention": [{"tokens": "Final pathology for case 2 was primary tumor size 7.6 cm , RCC , clear cell type , Fuhrman grade 3 , stage T3bNxM1 , with negative margins ( Figure 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 29, "end_tok": 31}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 18, "image": "20465800_F8.jpg", "pair_id": "20465800_F8", "text": "What is the image showing?\n", "gpt4_answer": "The image is likely a graphical representation of the classification of 47 infants with biliary atresia into groups based on inflammation or fibrosis. The classification is done using differential histological scores or molecular profiling at the time of diagnosis.", "fig_caption": "Classification of 47 infants with biliary atresia into groups of inflammation or fibrosis based on differential histological scores \u22651 or \u22652 or on molecular profiling at diagnosis.", "in_text_mention": [{"tokens": "From the entire cohort of 47 subjects , 14 ( 30 % ) had a differential score of \u22651 for inflammation and 17 ( 36 % ) for fibrosis ; the remaining 16 ( 34 % ) were unclassified due to the differences between inflammation and fibrosis being zero ( Figure 8 ) .", "entities": [{"type": "figure", "entity_id": "8", "start_tok": 50, "end_tok": 52}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 8"}
-{"question_id": 19, "image": "20465800_F8.jpg", "pair_id": "20465800_F8", "text": "How many infants had a differential score of \u22651 for inflammation?\n", "gpt4_answer": "There were 14 infants with a differential score of \u22651 for inflammation.", "fig_caption": "Classification of 47 infants with biliary atresia into groups of inflammation or fibrosis based on differential histological scores \u22651 or \u22652 or on molecular profiling at diagnosis.", "in_text_mention": [{"tokens": "From the entire cohort of 47 subjects , 14 ( 30 % ) had a differential score of \u22651 for inflammation and 17 ( 36 % ) for fibrosis ; the remaining 16 ( 34 % ) were unclassified due to the differences between inflammation and fibrosis being zero ( Figure 8 ) .", "entities": [{"type": "figure", "entity_id": "8", "start_tok": 50, "end_tok": 52}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 8"}
-{"question_id": 20, "image": "20465800_F8.jpg", "pair_id": "20465800_F8", "text": "How many infants had a differential score of \u22652 for fibrosis?\n", "gpt4_answer": "There were 17 infants with a differential score of \u22652 for fibrosis.", "fig_caption": "Classification of 47 infants with biliary atresia into groups of inflammation or fibrosis based on differential histological scores \u22651 or \u22652 or on molecular profiling at diagnosis.", "in_text_mention": [{"tokens": "From the entire cohort of 47 subjects , 14 ( 30 % ) had a differential score of \u22651 for inflammation and 17 ( 36 % ) for fibrosis ; the remaining 16 ( 34 % ) were unclassified due to the differences between inflammation and fibrosis being zero ( Figure 8 ) .", "entities": [{"type": "figure", "entity_id": "8", "start_tok": 50, "end_tok": 52}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 8"}
-{"question_id": 21, "image": "20671840_F0004.jpg", "pair_id": "20671840_F0004", "text": "What is the main finding in the orbital MRI?\n", "gpt4_answer": "The main finding in the orbital MRI is bilateral advanced proptosis, which is the abnormal protrusion of the eyeballs. Additionally, there is diffuse infiltration of orbital fat, obliteration of optic nerves, extraocular muscles with fixation of intraorbital structures, and enlarged lacrimal glands. The extraocular muscles show diffuse enlargement, including both tendons and muscle bundles, in a tubular configuration. There is also involvement of the perineural optic sheath.", "fig_caption": "The orbital MRI reveals bilateral advanced proptosis with diffuse infiltration of orbital fat, obliteration of optic nerves, extraocular muscles with fi xation of intraorbital structures and enlarged lacrimal glands. Extraocular muscles show diffuse enlargement (both tendons as well muscle bundles enlarged in a tubular confi guration), with involvement of perineural optic sheath", "in_text_mention": [{"tokens": "The extraocular muscles were diffusely enlarged ( tendons as well as muscle bellies in a tubular configuration ) [ Figure 4 ] .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 22, "image": "20671840_F0004.jpg", "pair_id": "20671840_F0004", "text": "What does proptosis mean?\n", "gpt4_answer": "Proptosis, also known as exophthalmos, refers to the abnormal protrusion or bulging of one or both eyeballs. It can be caused by various factors, such as inflammation, infection, injury, or tumors. In some cases, proptosis can be a symptom of an underlying medical condition, such as thyroid eye disease (Graves' ophthalmopathy) or orbital cellulitis. The severity of proptosis can range from mild to severe, and it may affect vision, eye movement, and the appearance of the eyes.", "fig_caption": "The orbital MRI reveals bilateral advanced proptosis with diffuse infiltration of orbital fat, obliteration of optic nerves, extraocular muscles with fi xation of intraorbital structures and enlarged lacrimal glands. Extraocular muscles show diffuse enlargement (both tendons as well muscle bundles enlarged in a tubular confi guration), with involvement of perineural optic sheath", "in_text_mention": [{"tokens": "The extraocular muscles were diffusely enlarged ( tendons as well as muscle bellies in a tubular configuration ) [ Figure 4 ] .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 23, "image": "20671840_F0004.jpg", "pair_id": "20671840_F0004", "text": "What can you say about the extraocular muscles?\n", "gpt4_answer": "In the orbital MRI, the extraocular muscles show diffuse enlargement, which affects both the tendons and muscle bundles in a tubular configuration. The extraocular muscles are responsible for controlling eye movement, and their enlargement may be related to the observed proptosis and other abnormalities in the orbital region. The cause of the enlargement could be due to inflammation, infection, or other pathological processes. Further evaluation and clinical correlation are needed to determine the underlying cause and appropriate treatment for these findings.", "fig_caption": "The orbital MRI reveals bilateral advanced proptosis with diffuse infiltration of orbital fat, obliteration of optic nerves, extraocular muscles with fi xation of intraorbital structures and enlarged lacrimal glands. Extraocular muscles show diffuse enlargement (both tendons as well muscle bundles enlarged in a tubular confi guration), with involvement of perineural optic sheath", "in_text_mention": [{"tokens": "The extraocular muscles were diffusely enlarged ( tendons as well as muscle bellies in a tubular configuration ) [ Figure 4 ] .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 24, "image": "20733967_F3.jpg", "pair_id": "20733967_F3", "text": "What is the main finding in the image?\n", "gpt4_answer": "The main finding in the image is an aneurysm of the left internal iliac artery, which is protruding into the urinary bladder.", "fig_caption": "Contrast-enhanced pelvic CT scan showing the aneurysm of the left internal iliac artery protruding into the urinary bladder (transverse view).", "in_text_mention": [{"tokens": "The greatest diameter of the aortic aneurysm was 8.0 cm , and the left internal iliac artery 's diameter was 11.5 cm ( Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 23, "end_tok": 25}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "FIG. 3"}
-{"question_id": 25, "image": "20733967_F3.jpg", "pair_id": "20733967_F3", "text": "What type of scan is it?\n", "gpt4_answer": "It is a contrast-enhanced pelvic CT (computed tomography) scan.", "fig_caption": "Contrast-enhanced pelvic CT scan showing the aneurysm of the left internal iliac artery protruding into the urinary bladder (transverse view).", "in_text_mention": [{"tokens": "The greatest diameter of the aortic aneurysm was 8.0 cm , and the left internal iliac artery 's diameter was 11.5 cm ( Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 23, "end_tok": 25}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "FIG. 3"}
-{"question_id": 26, "image": "20733967_F3.jpg", "pair_id": "20733967_F3", "text": "Is the aneurysm in the aortic aneurysm or the left internal iliac artery?\n", "gpt4_answer": "The aneurysm is in the left internal iliac artery.", "fig_caption": "Contrast-enhanced pelvic CT scan showing the aneurysm of the left internal iliac artery protruding into the urinary bladder (transverse view).", "in_text_mention": [{"tokens": "The greatest diameter of the aortic aneurysm was 8.0 cm , and the left internal iliac artery 's diameter was 11.5 cm ( Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 23, "end_tok": 25}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "FIG. 3"}
-{"question_id": 27, "image": "21139713_F0003.jpg", "pair_id": "21139713_F0003", "text": "What procedure was performed on the patient?\n", "gpt4_answer": "The patient underwent a thoracoplasty, which is a surgical procedure that involves the removal or reshaping of ribs to collapse a part of the chest cavity. This procedure is typically performed to treat conditions such as chronic lung infections or persistent air leaks in the chest cavity.", "fig_caption": "Chest X ray after thoracoplasty", "in_text_mention": [{"tokens": "Of the 16 patients in the tuberculous empyema group who needed surgery , 4 required decortication only , 10 patients required decortication with closure of bronchochopleural fistula [ Figure 2 ] , and 2 of them needed additional thoracoplasty [ Figure 3 ] .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 40, "end_tok": 42}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 28, "image": "21139713_F0003.jpg", "pair_id": "21139713_F0003", "text": "What was the purpose of the thoracoplasty in this case?\n", "gpt4_answer": "In this specific case, the thoracoplasty was performed as an additional procedure for a patient with tuberculous empyema. Tuberculous empyema is a condition where pus accumulates in the pleural space due to a tuberculosis infection. The thoracoplasty was likely performed to help manage the infection and improve the patient's condition by collapsing a part of the chest cavity.", "fig_caption": "Chest X ray after thoracoplasty", "in_text_mention": [{"tokens": "Of the 16 patients in the tuberculous empyema group who needed surgery , 4 required decortication only , 10 patients required decortication with closure of bronchochopleural fistula [ Figure 2 ] , and 2 of them needed additional thoracoplasty [ Figure 3 ] .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 40, "end_tok": 42}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 29, "image": "22174521_F2.jpg", "pair_id": "22174521_F2", "text": "What is the organ in the image?\n", "gpt4_answer": "The organ in the image is the right kidney.", "fig_caption": "Gross section of Right kidney showing xanthogranulomatous area (arrow)", "in_text_mention": [{"tokens": "The cut section of the gross specimen showed xanthomatous area [ Figure 2 ] and the histopathology was suggestive of XPN [ Figure 3 ] .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 11, "end_tok": 13}]}, {"tokens": "XPN was subsequently established on evaluation of gross specimen [ Figure 2 ] and histopathology [ Figure 3 ] .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 10, "end_tok": 12}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 30, "image": "22174521_F2.jpg", "pair_id": "22174521_F2", "text": "What are the arrows pointing to?\n", "gpt4_answer": "The arrows in the image are pointing to a xanthogranulomatous area within the right kidney.", "fig_caption": "Gross section of Right kidney showing xanthogranulomatous area (arrow)", "in_text_mention": [{"tokens": "The cut section of the gross specimen showed xanthomatous area [ Figure 2 ] and the histopathology was suggestive of XPN [ Figure 3 ] .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 11, "end_tok": 13}]}, {"tokens": "XPN was subsequently established on evaluation of gross specimen [ Figure 2 ] and histopathology [ Figure 3 ] .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 10, "end_tok": 12}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 31, "image": "22174521_F2.jpg", "pair_id": "22174521_F2", "text": "What does xanthogranulomatous mean?\n", "gpt4_answer": "Xanthogranulomatous refers to a type of chronic inflammation characterized by the presence of lipid-laden macrophages, which are immune cells that have ingested lipids (fats). This type of inflammation can occur in various organs, but in the context of the image, it is seen in the kidney. Xanthogranulomatous inflammation can lead to the destruction of the normal tissue architecture and replacement with granulomatous tissue, which consists of immune cells and fibrosis (scar tissue). The cause of xanthogranulomatous inflammation can vary, but it is often associated with long-standing infections or other chronic inflammatory conditions.", "fig_caption": "Gross section of Right kidney showing xanthogranulomatous area (arrow)", "in_text_mention": [{"tokens": "The cut section of the gross specimen showed xanthomatous area [ Figure 2 ] and the histopathology was suggestive of XPN [ Figure 3 ] .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 11, "end_tok": 13}]}, {"tokens": "XPN was subsequently established on evaluation of gross specimen [ Figure 2 ] and histopathology [ Figure 3 ] .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 10, "end_tok": 12}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 32, "image": "22223938_F5.jpg", "pair_id": "22223938_F5", "text": "What is the color of the mass?\n", "gpt4_answer": "The color of the mass in the image is brownish-yellow.", "fig_caption": "Cut surface of the gross specimen shows the brownish-yellow color of the mass, with multiple vessels but without any areas of hemorrhage or necrosis", "in_text_mention": [{"tokens": "The cut surface was brownish yellow in color , with multiple coursing vessels and without any areas of hemorrhage or necrosis [ Figure 5 ] .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 22, "end_tok": 24}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 33, "image": "22223938_F5.jpg", "pair_id": "22223938_F5", "text": "Are there any areas of hemorrhage or necrosis?\n", "gpt4_answer": "No, there are no areas of hemorrhage or necrosis visible in the image.", "fig_caption": "Cut surface of the gross specimen shows the brownish-yellow color of the mass, with multiple vessels but without any areas of hemorrhage or necrosis", "in_text_mention": [{"tokens": "The cut surface was brownish yellow in color , with multiple coursing vessels and without any areas of hemorrhage or necrosis [ Figure 5 ] .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 22, "end_tok": 24}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 34, "image": "22223938_F5.jpg", "pair_id": "22223938_F5", "text": "What can you tell me about the vessels?\n", "gpt4_answer": "The image shows multiple coursing vessels within the mass. This suggests that the mass has a vascular component, meaning it has a network of blood vessels supplying it. The presence of vessels can be an important factor when considering the nature of the mass and its potential for growth or spread. However, further analysis and clinical correlation are needed to determine the cause and significance of these findings.", "fig_caption": "Cut surface of the gross specimen shows the brownish-yellow color of the mass, with multiple vessels but without any areas of hemorrhage or necrosis", "in_text_mention": [{"tokens": "The cut surface was brownish yellow in color , with multiple coursing vessels and without any areas of hemorrhage or necrosis [ Figure 5 ] .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 22, "end_tok": 24}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 35, "image": "22529899_pone-0033860-g004.jpg", "pair_id": "22529899_pone-0033860-g004", "text": "What are the major tissues obtained from the mice?\n", "gpt4_answer": "The major tissues obtained from the mice include the heart, liver, spleen, lung, and kidney.", "fig_caption": "\nSystematic toxicity in the SAHA-DDP/PECE treated mice. H&E staining of section of major tissues obtained from tumor beard mice which received two doses of NS, PECE, SAHA, DDP, SAHA-DDP and SAHA-DDP/PECE. Heart, liver, spleen, lung and kidney were harvested at day 28 after intravenous injection. (A) Mean body weights on days 7, 10, 13, 16, 19, 22, 25 and 28 of mice treated with two doses of NS, PECE, SAHA, DDP, SAHA-DDP and SAHA-DDP/PECE; error bars correspond to 95% confidence intervals, Values are means\u00b1SD (n \u200a=\u200a 10 mice per group).", "in_text_mention": [{"tokens": "No difference of body weight was observed among the SAHA-DDP / PECE group compared with the other groups at the end of the study ( Fig. 4A ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 25, "end_tok": 27}]}, {"tokens": "As observed by two pathologists in a blinded manner , we found that mice treated with SAHA-DDP / PECE showed no slight changes of toxicity to the organ tissues , compared with the normal organ tissues from mice receiving NS ( Fig. 4B ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 41, "end_tok": 43}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 36, "image": "22529899_pone-0033860-g004.jpg", "pair_id": "22529899_pone-0033860-g004", "text": "What is the purpose of H&E staining?\n", "gpt4_answer": "H&E staining, or hematoxylin and eosin staining, is a widely used technique in histology to visualize the structure of cells and tissues in biological samples. Hematoxylin stains the cell nuclei blue, while eosin stains the cytoplasm and extracellular matrix pink. This staining method helps to differentiate various cell types and tissue structures, allowing researchers and pathologists to examine the morphology and organization of cells within the tissue, as well as to identify any abnormalities or pathological changes.", "fig_caption": "\nSystematic toxicity in the SAHA-DDP/PECE treated mice. H&E staining of section of major tissues obtained from tumor beard mice which received two doses of NS, PECE, SAHA, DDP, SAHA-DDP and SAHA-DDP/PECE. Heart, liver, spleen, lung and kidney were harvested at day 28 after intravenous injection. (A) Mean body weights on days 7, 10, 13, 16, 19, 22, 25 and 28 of mice treated with two doses of NS, PECE, SAHA, DDP, SAHA-DDP and SAHA-DDP/PECE; error bars correspond to 95% confidence intervals, Values are means\u00b1SD (n \u200a=\u200a 10 mice per group).", "in_text_mention": [{"tokens": "No difference of body weight was observed among the SAHA-DDP / PECE group compared with the other groups at the end of the study ( Fig. 4A ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 25, "end_tok": 27}]}, {"tokens": "As observed by two pathologists in a blinded manner , we found that mice treated with SAHA-DDP / PECE showed no slight changes of toxicity to the organ tissues , compared with the normal organ tissues from mice receiving NS ( Fig. 4B ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 41, "end_tok": 43}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 37, "image": "22529899_pone-0033860-g004.jpg", "pair_id": "22529899_pone-0033860-g004", "text": "What is the significance of the mean body weights in the study?\n", "gpt4_answer": "The mean body weights of the mice in the study are used to assess the potential systemic toxicity of the treatments being tested. By monitoring the body weights of the mice over time, researchers can determine if any of the treatments are causing significant weight loss or gain, which could indicate adverse effects on the overall health of the mice. In this case, the mean body weights were measured on various days after the treatments were administered, and the error bars correspond to 95% confidence intervals. Comparing the body weights among different treatment groups can help researchers evaluate the safety and tolerability of the treatments being studied.", "fig_caption": "\nSystematic toxicity in the SAHA-DDP/PECE treated mice. H&E staining of section of major tissues obtained from tumor beard mice which received two doses of NS, PECE, SAHA, DDP, SAHA-DDP and SAHA-DDP/PECE. Heart, liver, spleen, lung and kidney were harvested at day 28 after intravenous injection. (A) Mean body weights on days 7, 10, 13, 16, 19, 22, 25 and 28 of mice treated with two doses of NS, PECE, SAHA, DDP, SAHA-DDP and SAHA-DDP/PECE; error bars correspond to 95% confidence intervals, Values are means\u00b1SD (n \u200a=\u200a 10 mice per group).", "in_text_mention": [{"tokens": "No difference of body weight was observed among the SAHA-DDP / PECE group compared with the other groups at the end of the study ( Fig. 4A ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 25, "end_tok": 27}]}, {"tokens": "As observed by two pathologists in a blinded manner , we found that mice treated with SAHA-DDP / PECE showed no slight changes of toxicity to the organ tissues , compared with the normal organ tissues from mice receiving NS ( Fig. 4B ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 41, "end_tok": 43}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 38, "image": "22569536_fig2.jpg", "pair_id": "22569536_fig2", "text": "What is the main focus of the figure?\n", "gpt4_answer": "The main focus of the figure is to show the overall survival of patients according to Lauren's histological type of carcinoma, with or without HER2 gene amplification. The data is likely presented in the form of a survival curve or a similar graphical representation, which helps visualize the relationship between the histological type of carcinoma, HER2 gene amplification status, and patient survival over time.", "fig_caption": "Overall survival according to Lauren\u2019s histological type of carcinoma with or without HER2 gene amplification by dual colour silver enhanced in situ hybridisation.", "in_text_mention": [{"tokens": "The Cox regression analysis model showed similar results ( p = 0.0005 ) ( figure 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 14, "end_tok": 16}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 39, "image": "22569536_fig2.jpg", "pair_id": "22569536_fig2", "text": "What is the method used to determine HER2 gene amplification?\n", "gpt4_answer": "The method used to determine HER2 gene amplification in this study is dual-color silver-enhanced in situ hybridization (SISH). This technique allows for the visualization of specific DNA sequences within cells, in this case, the HER2 gene, and helps to determine if there is an increased number of copies of the gene, which is referred to as gene amplification.", "fig_caption": "Overall survival according to Lauren\u2019s histological type of carcinoma with or without HER2 gene amplification by dual colour silver enhanced in situ hybridisation.", "in_text_mention": [{"tokens": "The Cox regression analysis model showed similar results ( p = 0.0005 ) ( figure 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 14, "end_tok": 16}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 40, "image": "22569536_fig2.jpg", "pair_id": "22569536_fig2", "text": "What is the significance of the Cox regression analysis model in this context?\n", "gpt4_answer": "In this context, the Cox regression analysis model is used to assess the relationship between the overall survival of patients and various factors, such as Lauren's histological type of carcinoma and HER2 gene amplification status. The model helps to determine if these factors have a statistically significant impact on patient survival. The p-value mentioned (p = 0.0005) suggests that there is a statistically significant association between the factors being analyzed and overall survival.", "fig_caption": "Overall survival according to Lauren\u2019s histological type of carcinoma with or without HER2 gene amplification by dual colour silver enhanced in situ hybridisation.", "in_text_mention": [{"tokens": "The Cox regression analysis model showed similar results ( p = 0.0005 ) ( figure 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 14, "end_tok": 16}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 41, "image": "23493967_F1.jpg", "pair_id": "23493967_F1", "text": "What does the chest X-ray show?\n", "gpt4_answer": "The chest X-ray shows dilated cardiomyopathy, which is a condition where the heart becomes enlarged and weakened, affecting its ability to pump blood effectively.", "fig_caption": "Chest X-ray showing dilated cardiomyopathy", "in_text_mention": [{"tokens": "Severe mitral regurgitation was also diagnosed , and by 3 years , he developed dilated cardiomyopathy ( DCM ) [ Figure 1 ] with heart failure .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 20, "end_tok": 22}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 42, "image": "23493967_F1.jpg", "pair_id": "23493967_F1", "text": "What is the condition of the heart?\n", "gpt4_answer": "The heart appears to be enlarged and weakened, which is characteristic of dilated cardiomyopathy. This condition can lead to a decreased ability of the heart to pump blood effectively, and may result in heart failure.", "fig_caption": "Chest X-ray showing dilated cardiomyopathy", "in_text_mention": [{"tokens": "Severe mitral regurgitation was also diagnosed , and by 3 years , he developed dilated cardiomyopathy ( DCM ) [ Figure 1 ] with heart failure .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 20, "end_tok": 22}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 43, "image": "23493967_F1.jpg", "pair_id": "23493967_F1", "text": "Is there any other condition mentioned in the context?\n", "gpt4_answer": "Yes, there is another condition mentioned, which is severe mitral regurgitation. Mitral regurgitation is a condition where the mitral valve in the heart does not close properly, allowing blood to flow backward into the left atrium when the left ventricle contracts. This can lead to a decrease in the efficiency of the heart's pumping action and may contribute to the development of heart failure.", "fig_caption": "Chest X-ray showing dilated cardiomyopathy", "in_text_mention": [{"tokens": "Severe mitral regurgitation was also diagnosed , and by 3 years , he developed dilated cardiomyopathy ( DCM ) [ Figure 1 ] with heart failure .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 20, "end_tok": 22}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 44, "image": "23565458_F1.jpg", "pair_id": "23565458_F1", "text": "What do the opacities look like?\n", "gpt4_answer": "The opacities in the chest X-ray appear as fluffy or hazy areas in both lung fields. These opacities can be indicative of various lung conditions, such as infections, inflammation, or other lung abnormalities.", "fig_caption": "Chest X-ray showing pluffy opacities in both lung fields", "in_text_mention": null, "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 45, "image": "23565458_F1.jpg", "pair_id": "23565458_F1", "text": "Are the opacities present in both lungs?\n", "gpt4_answer": "Yes, the opacities are present in both lung fields, which means they are affecting both the left and right lungs.", "fig_caption": "Chest X-ray showing pluffy opacities in both lung fields", "in_text_mention": null, "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 46, "image": "23565458_F1.jpg", "pair_id": "23565458_F1", "text": "What could be the possible cause of these opacities?\n", "gpt4_answer": "There are several possible causes for the fluffy opacities seen in both lung fields. Some common causes include infections (such as pneumonia), inflammation, fluid accumulation, or other lung abnormalities. It is important to consider the patient's clinical history, symptoms, and additional diagnostic tests to determine the exact cause and appropriate treatment. A healthcare professional should be consulted for a thorough evaluation and proper diagnosis.", "fig_caption": "Chest X-ray showing pluffy opacities in both lung fields", "in_text_mention": null, "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 47, "image": "24179348_f4c-ccrep-1-2008-057.jpg", "pair_id": "24179348_f4c-ccrep-1-2008-057", "text": "What is being compressed in the image?\n", "gpt4_answer": "In the image, the esophagus is being compressed.", "fig_caption": "Sagittal T2-w MRI demonstrates compression of the esophagus by anterior osteophytes.", "in_text_mention": [{"tokens": "The length of styloid process were elongated on both sides ( right : 29 mm , left : 30 mm ) ( Fig. 4a ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 22, "end_tok": 24}]}, {"tokens": "Barium swallow test revealed indentations along the esophagus , which were more prominent at C5-6 and C6-7 ( Fig. 4b ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 18, "end_tok": 20}]}, {"tokens": "Cervical MRI showed anterior osteophyte formations at the same levels which compressed the esophagus ( Fig. 4c ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 15, "end_tok": 17}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 4c"}
-{"question_id": 48, "image": "24179348_f4c-ccrep-1-2008-057.jpg", "pair_id": "24179348_f4c-ccrep-1-2008-057", "text": "What is causing the compression?\n", "gpt4_answer": "The compression of the esophagus is caused by anterior osteophytes.", "fig_caption": "Sagittal T2-w MRI demonstrates compression of the esophagus by anterior osteophytes.", "in_text_mention": [{"tokens": "The length of styloid process were elongated on both sides ( right : 29 mm , left : 30 mm ) ( Fig. 4a ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 22, "end_tok": 24}]}, {"tokens": "Barium swallow test revealed indentations along the esophagus , which were more prominent at C5-6 and C6-7 ( Fig. 4b ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 18, "end_tok": 20}]}, {"tokens": "Cervical MRI showed anterior osteophyte formations at the same levels which compressed the esophagus ( Fig. 4c ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 15, "end_tok": 17}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 4c"}
-{"question_id": 49, "image": "24179348_f4c-ccrep-1-2008-057.jpg", "pair_id": "24179348_f4c-ccrep-1-2008-057", "text": "What type of MRI is used in this image?\n", "gpt4_answer": "The image is a sagittal T2-weighted MRI.", "fig_caption": "Sagittal T2-w MRI demonstrates compression of the esophagus by anterior osteophytes.", "in_text_mention": [{"tokens": "The length of styloid process were elongated on both sides ( right : 29 mm , left : 30 mm ) ( Fig. 4a ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 22, "end_tok": 24}]}, {"tokens": "Barium swallow test revealed indentations along the esophagus , which were more prominent at C5-6 and C6-7 ( Fig. 4b ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 18, "end_tok": 20}]}, {"tokens": "Cervical MRI showed anterior osteophyte formations at the same levels which compressed the esophagus ( Fig. 4c ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 15, "end_tok": 17}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 4c"}
-{"question_id": 50, "image": "24554380_Fig23.jpg", "pair_id": "24554380_Fig23", "text": "What type of fracture is shown in the image?\n", "gpt4_answer": "The image shows an avulsion fracture in the context of a hyperextension dislocation.", "fig_caption": "Avulsion fracture in hyperextension dislocation. Sagittal CT image shows an avulsion fracture at the anterosuperior end plate of C7 (white arrow), with the transverse diameter wider than the vertical diameter. Avulsion of the posteroinferior corner of the C6 vertebral body is also seen (black arrow), which suggests avulsion of the posterior annulus or PLL", "in_text_mention": [{"tokens": "Less common CT indicators include : disc vacuum phenomenon and an avulsion fracture of the anteroinferior margin of the involved vertebra caused by avulsion due to the intact Sharpey\u2019s fibres ( Fig. 23 ) [ 23 ] .", "entities": [{"type": "figure", "entity_id": "23", "start_tok": 31, "end_tok": 33}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig. 23"}
-{"question_id": 51, "image": "24554380_Fig23.jpg", "pair_id": "24554380_Fig23", "text": "Which vertebrae are affected?\n", "gpt4_answer": "The affected vertebrae are C7 and C6, which are part of the cervical spine.", "fig_caption": "Avulsion fracture in hyperextension dislocation. Sagittal CT image shows an avulsion fracture at the anterosuperior end plate of C7 (white arrow), with the transverse diameter wider than the vertical diameter. Avulsion of the posteroinferior corner of the C6 vertebral body is also seen (black arrow), which suggests avulsion of the posterior annulus or PLL", "in_text_mention": [{"tokens": "Less common CT indicators include : disc vacuum phenomenon and an avulsion fracture of the anteroinferior margin of the involved vertebra caused by avulsion due to the intact Sharpey\u2019s fibres ( Fig. 23 ) [ 23 ] .", "entities": [{"type": "figure", "entity_id": "23", "start_tok": 31, "end_tok": 33}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig. 23"}
-{"question_id": 52, "image": "24554380_Fig23.jpg", "pair_id": "24554380_Fig23", "text": "What are the arrows pointing to?\n", "gpt4_answer": "The white arrow is pointing to an avulsion fracture at the anterosuperior end plate of the C7 vertebra, while the black arrow is pointing to an avulsion of the posteroinferior corner of the C6 vertebral body. These findings suggest that there may be an avulsion of the posterior annulus or the posterior longitudinal ligament (PLL).", "fig_caption": "Avulsion fracture in hyperextension dislocation. Sagittal CT image shows an avulsion fracture at the anterosuperior end plate of C7 (white arrow), with the transverse diameter wider than the vertical diameter. Avulsion of the posteroinferior corner of the C6 vertebral body is also seen (black arrow), which suggests avulsion of the posterior annulus or PLL", "in_text_mention": [{"tokens": "Less common CT indicators include : disc vacuum phenomenon and an avulsion fracture of the anteroinferior margin of the involved vertebra caused by avulsion due to the intact Sharpey\u2019s fibres ( Fig. 23 ) [ 23 ] .", "entities": [{"type": "figure", "entity_id": "23", "start_tok": 31, "end_tok": 33}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig. 23"}
-{"question_id": 53, "image": "24614132_pone-0091649-g004.jpg", "pair_id": "24614132_pone-0091649-g004", "text": "What is the scatter matrix showing?\n", "gpt4_answer": "The scatter matrix in the image is a visual representation of the relationships between different PET/CT and DCE-MRI metrics in adenocarcinoma (AC) of the lung. It displays the correlations between various pairs of metrics, allowing for a quick assessment of the strength and direction of the relationships between them. Each cell in the matrix represents a scatter plot of two metrics, and the diagonal cells usually show the distribution of a single metric. The scatter matrix can help researchers identify patterns, trends, and potential outliers in the data.", "fig_caption": "Scatter matrix of the PET/CT and DCE-MRI metrics in AC of the lung.", "in_text_mention": [{"tokens": "3 ) , AC and SCC at a high level of statistical significance ( \u03c1 = 0.926 , P < 0.001 , Fig. 4 ; \u03c1 = 0.867 , P < 0.001 and \u03c1 = 0.968 , P < 0.001 , Fig. 5 , respectively ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 22, "end_tok": 24}]}, {"tokens": "3 ) ; for Ktrans and Kep ( \u03c1 = 0.606 , P = 0.001 ) , Ktrans and Ve ( \u03c1 = 0.537 , P = 0.006 ) , Ktrans and iAUC ( \u03c1 = 0.962 , P < 0.001 ) , Kep and iAUC ( \u03c1 = 0.566 , P = 0.003 ) and Ve and iAUC ( \u03c1 = 0.530 , P = 0.006 ) in AC ( Fig. 4 ) ; and for Ktrans and Ve ( \u03c1 = 0.776 , P < 0.001 ) , Ktrans and iAUC ( \u03c1 = 0.979 , P < 0.001 ) , and Ve and iAUC ( \u03c1 = 0.738 , P = 0.001 ) in SCC ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 71, "end_tok": 73}]}, {"tokens": "3 ) ; for SUVmax and iAUC ( \u03c1 = \u22120.420 , P = 0.037 ) , SUVmean and Ktrans ( \u03c1 = \u22120.411 , P = 0.041 ) , SUVmean and Kep ( \u03c1 = \u22120.045 , P = 0.026 ) , and SUVmean and iAUC ( \u03c1 = \u22120.512 , P = 0.009 ) in AC ( Fig. 4 ) ; However , for neither in SCC ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 59, "end_tok": 61}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 54, "image": "24614132_pone-0091649-g004.jpg", "pair_id": "24614132_pone-0091649-g004", "text": "What are PET/CT and DCE-MRI?\n", "gpt4_answer": "PET/CT and DCE-MRI are advanced medical imaging techniques used to visualize and assess various aspects of the body's internal structures and functions.\n\nPET/CT (Positron Emission Tomography/Computed Tomography) is a combination of two imaging techniques: PET, which uses a small amount of radioactive material to visualize metabolic processes in the body, and CT, which uses X-rays to create detailed cross-sectional images of the body's structures. PET/CT is often used to detect and monitor cancer, as well as to evaluate the effectiveness of cancer treatments.\n\nDCE-MRI (Dynamic Contrast-Enhanced Magnetic Resonance Imaging) is a specialized form of MRI that involves the injection of a contrast agent into the bloodstream. This technique allows for the visualization of blood flow and vascular permeability in tissues, which can be useful in assessing tumors and other abnormalities. DCE-MRI is often used to evaluate the response to cancer treatments and to monitor the progression of diseases.", "fig_caption": "Scatter matrix of the PET/CT and DCE-MRI metrics in AC of the lung.", "in_text_mention": [{"tokens": "3 ) , AC and SCC at a high level of statistical significance ( \u03c1 = 0.926 , P < 0.001 , Fig. 4 ; \u03c1 = 0.867 , P < 0.001 and \u03c1 = 0.968 , P < 0.001 , Fig. 5 , respectively ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 22, "end_tok": 24}]}, {"tokens": "3 ) ; for Ktrans and Kep ( \u03c1 = 0.606 , P = 0.001 ) , Ktrans and Ve ( \u03c1 = 0.537 , P = 0.006 ) , Ktrans and iAUC ( \u03c1 = 0.962 , P < 0.001 ) , Kep and iAUC ( \u03c1 = 0.566 , P = 0.003 ) and Ve and iAUC ( \u03c1 = 0.530 , P = 0.006 ) in AC ( Fig. 4 ) ; and for Ktrans and Ve ( \u03c1 = 0.776 , P < 0.001 ) , Ktrans and iAUC ( \u03c1 = 0.979 , P < 0.001 ) , and Ve and iAUC ( \u03c1 = 0.738 , P = 0.001 ) in SCC ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 71, "end_tok": 73}]}, {"tokens": "3 ) ; for SUVmax and iAUC ( \u03c1 = \u22120.420 , P = 0.037 ) , SUVmean and Ktrans ( \u03c1 = \u22120.411 , P = 0.041 ) , SUVmean and Kep ( \u03c1 = \u22120.045 , P = 0.026 ) , and SUVmean and iAUC ( \u03c1 = \u22120.512 , P = 0.009 ) in AC ( Fig. 4 ) ; However , for neither in SCC ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 59, "end_tok": 61}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 55, "image": "24614132_pone-0091649-g004.jpg", "pair_id": "24614132_pone-0091649-g004", "text": "What is the significance of the correlations in the scatter matrix?\n", "gpt4_answer": "The correlations in the scatter matrix can provide valuable insights into the relationships between different PET/CT and DCE-MRI metrics in adenocarcinoma (AC) of the lung. By examining the strength and direction of these correlations, researchers can identify potential associations between the imaging metrics and various aspects of the disease, such as tumor growth, response to treatment, or disease progression.\n\nA strong positive correlation between two metrics suggests that they tend to increase or decrease together, while a strong negative correlation indicates that one metric tends to increase as the other decreases. Weak or no correlation suggests that there is little or no relationship between the two metrics.\n\nUnderstanding these correlations can help researchers and clinicians make more informed decisions about patient care, such as selecting the most appropriate imaging techniques for monitoring disease progression or evaluating the effectiveness of treatments. Additionally, the correlations can provide insights into the underlying biological processes and mechanisms involved in the disease, which may contribute to the development of new therapeutic strategies.", "fig_caption": "Scatter matrix of the PET/CT and DCE-MRI metrics in AC of the lung.", "in_text_mention": [{"tokens": "3 ) , AC and SCC at a high level of statistical significance ( \u03c1 = 0.926 , P < 0.001 , Fig. 4 ; \u03c1 = 0.867 , P < 0.001 and \u03c1 = 0.968 , P < 0.001 , Fig. 5 , respectively ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 22, "end_tok": 24}]}, {"tokens": "3 ) ; for Ktrans and Kep ( \u03c1 = 0.606 , P = 0.001 ) , Ktrans and Ve ( \u03c1 = 0.537 , P = 0.006 ) , Ktrans and iAUC ( \u03c1 = 0.962 , P < 0.001 ) , Kep and iAUC ( \u03c1 = 0.566 , P = 0.003 ) and Ve and iAUC ( \u03c1 = 0.530 , P = 0.006 ) in AC ( Fig. 4 ) ; and for Ktrans and Ve ( \u03c1 = 0.776 , P < 0.001 ) , Ktrans and iAUC ( \u03c1 = 0.979 , P < 0.001 ) , and Ve and iAUC ( \u03c1 = 0.738 , P = 0.001 ) in SCC ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 71, "end_tok": 73}]}, {"tokens": "3 ) ; for SUVmax and iAUC ( \u03c1 = \u22120.420 , P = 0.037 ) , SUVmean and Ktrans ( \u03c1 = \u22120.411 , P = 0.041 ) , SUVmean and Kep ( \u03c1 = \u22120.045 , P = 0.026 ) , and SUVmean and iAUC ( \u03c1 = \u22120.512 , P = 0.009 ) in AC ( Fig. 4 ) ; However , for neither in SCC ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 59, "end_tok": 61}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 56, "image": "24926336_f2-etm-07-06-1521.jpg", "pair_id": "24926336_f2-etm-07-06-1521", "text": "What is the main finding in the left-middle ureter?\n", "gpt4_answer": "The main finding in the left-middle ureter is a nodular soft-tissue density area at the L3 level.", "fig_caption": "PET-CT scans revealed a nodular soft-tissue density area in the wall of the left-middle ureter at the L3 level, paraaortic and iliac lymphadenopathy and increased FDG uptake in the sternum and right ilium. PET-CT, positron emission tomography-computed tomography; FDG, fludeoxyglucose.", "in_text_mention": [{"tokens": "Positron emission tomography-computed tomography ( PET-CT ) revealed an area of nodular soft-tissue density in the wall of the left middle ureter at the L3 level ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 57, "image": "24926336_f2-etm-07-06-1521.jpg", "pair_id": "24926336_f2-etm-07-06-1521", "text": "What other findings are mentioned in the image?\n", "gpt4_answer": "The image also shows paraaortic and iliac lymphadenopathy, as well as increased FDG (fludeoxyglucose) uptake in the sternum and right ilium.", "fig_caption": "PET-CT scans revealed a nodular soft-tissue density area in the wall of the left-middle ureter at the L3 level, paraaortic and iliac lymphadenopathy and increased FDG uptake in the sternum and right ilium. PET-CT, positron emission tomography-computed tomography; FDG, fludeoxyglucose.", "in_text_mention": [{"tokens": "Positron emission tomography-computed tomography ( PET-CT ) revealed an area of nodular soft-tissue density in the wall of the left middle ureter at the L3 level ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 58, "image": "24926336_f2-etm-07-06-1521.jpg", "pair_id": "24926336_f2-etm-07-06-1521", "text": "What imaging technique was used?\n", "gpt4_answer": "The imaging technique used in this case is PET-CT, which stands for positron emission tomography-computed tomography. This is a combination of two imaging techniques that provide both functional and anatomical information about the body. PET scans use a small amount of a radioactive tracer, such as FDG, to detect areas of increased metabolic activity, while CT scans provide detailed images of the body's internal structures.", "fig_caption": "PET-CT scans revealed a nodular soft-tissue density area in the wall of the left-middle ureter at the L3 level, paraaortic and iliac lymphadenopathy and increased FDG uptake in the sternum and right ilium. PET-CT, positron emission tomography-computed tomography; FDG, fludeoxyglucose.", "in_text_mention": [{"tokens": "Positron emission tomography-computed tomography ( PET-CT ) revealed an area of nodular soft-tissue density in the wall of the left middle ureter at the L3 level ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 59, "image": "27679760_fig2.jpg", "pair_id": "27679760_fig2", "text": "What organ is shown in the image?\n", "gpt4_answer": "The image shows a gross specimen of splenic tissue.", "fig_caption": "Gross specimen, with splenic tissue being darker in color.", "in_text_mention": [{"tokens": "The splenic tissue had a well delineated fibrous encapsulation , separate from the testicular tissue ( Figure 2 , Figure 3 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 16, "end_tok": 18}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure\u00a02"}
-{"question_id": 60, "image": "27679760_fig2.jpg", "pair_id": "27679760_fig2", "text": "What is the color of the splenic tissue?\n", "gpt4_answer": "The splenic tissue appears to be darker in color compared to the surrounding tissue.", "fig_caption": "Gross specimen, with splenic tissue being darker in color.", "in_text_mention": [{"tokens": "The splenic tissue had a well delineated fibrous encapsulation , separate from the testicular tissue ( Figure 2 , Figure 3 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 16, "end_tok": 18}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure\u00a02"}
-{"question_id": 61, "image": "27679760_fig2.jpg", "pair_id": "27679760_fig2", "text": "Is the splenic tissue separate from the testicular tissue?\n", "gpt4_answer": "Yes, the splenic tissue is separate from the testicular tissue, as it has a well-delineated fibrous encapsulation. This encapsulation creates a distinct boundary between the splenic and testicular tissues.", "fig_caption": "Gross specimen, with splenic tissue being darker in color.", "in_text_mention": [{"tokens": "The splenic tissue had a well delineated fibrous encapsulation , separate from the testicular tissue ( Figure 2 , Figure 3 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 16, "end_tok": 18}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure\u00a02"}
-{"question_id": 62, "image": "28399828_Fig5.jpg", "pair_id": "28399828_Fig5", "text": "What type of tissue is shown in the image?\n", "gpt4_answer": "The image shows a histopathological section of a disc fragment, which is a part of the intervertebral disc in the spine.", "fig_caption": "Histopathological findings. Representative images of HE-stained sections of the disc fragment extracted during the operation, showing liquefaction degeneration and granulation surrounding the sequestered disc fragment. Scale bar: 200\u00a0\u03bcm (hematoxylin and eosin [H & E]\u2009\u00d7\u200940)", "in_text_mention": [{"tokens": "Consistent with observations during the operation , histopathological analysis revealed liquefaction degeneration and granulation surrounding the sequestered disc fragment ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 20, "end_tok": 22}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Fig. 5"}
-{"question_id": 63, "image": "28399828_Fig5.jpg", "pair_id": "28399828_Fig5", "text": "What is the staining technique used?\n", "gpt4_answer": "The staining technique used in this image is hematoxylin and eosin (H&E) staining.", "fig_caption": "Histopathological findings. Representative images of HE-stained sections of the disc fragment extracted during the operation, showing liquefaction degeneration and granulation surrounding the sequestered disc fragment. Scale bar: 200\u00a0\u03bcm (hematoxylin and eosin [H & E]\u2009\u00d7\u200940)", "in_text_mention": [{"tokens": "Consistent with observations during the operation , histopathological analysis revealed liquefaction degeneration and granulation surrounding the sequestered disc fragment ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 20, "end_tok": 22}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Fig. 5"}
-{"question_id": 64, "image": "28399828_Fig5.jpg", "pair_id": "28399828_Fig5", "text": "What are the findings in the image?\n", "gpt4_answer": "The histopathological findings in the image show liquefaction degeneration and granulation surrounding the sequestered disc fragment. Liquefaction degeneration refers to the breakdown and disintegration of the tissue, while granulation refers to the formation of new, small blood vessels and connective tissue in response to injury or inflammation. These findings suggest that there is a pathological process affecting the intervertebral disc, which may be related to disc herniation or other spinal conditions.", "fig_caption": "Histopathological findings. Representative images of HE-stained sections of the disc fragment extracted during the operation, showing liquefaction degeneration and granulation surrounding the sequestered disc fragment. Scale bar: 200\u00a0\u03bcm (hematoxylin and eosin [H & E]\u2009\u00d7\u200940)", "in_text_mention": [{"tokens": "Consistent with observations during the operation , histopathological analysis revealed liquefaction degeneration and granulation surrounding the sequestered disc fragment ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 20, "end_tok": 22}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Fig. 5"}
-{"question_id": 65, "image": "29201784_F3.jpg", "pair_id": "29201784_F3", "text": "What are the main findings in the liver biopsy?\n", "gpt4_answer": "The main findings in the liver biopsy include interphase activity, confluent necrosis, hepatic duct proliferation, and biliary stasis. These observations suggest that there may be some pathological changes occurring in the liver tissue.", "fig_caption": "Histopathological finding of the liver", "in_text_mention": [{"tokens": "In liver biopsy interphase activity , confluent necrosis , hepatic duct proliferation , and biliary stasis were detected ( Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Fig. 3:"}
-{"question_id": 66, "image": "29201784_F3.jpg", "pair_id": "29201784_F3", "text": "What does confluent necrosis mean?\n", "gpt4_answer": "Confluent necrosis refers to the widespread death of cells or tissues in a continuous or merging pattern. In the context of a liver biopsy, it indicates that there is a significant amount of cell death occurring in the liver tissue, which may be due to various factors such as inflammation, infection, or other underlying conditions.", "fig_caption": "Histopathological finding of the liver", "in_text_mention": [{"tokens": "In liver biopsy interphase activity , confluent necrosis , hepatic duct proliferation , and biliary stasis were detected ( Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Fig. 3:"}
-{"question_id": 67, "image": "29201784_F3.jpg", "pair_id": "29201784_F3", "text": "What is biliary stasis?\n", "gpt4_answer": "Biliary stasis is a condition in which the flow of bile is slowed down or stopped within the liver or bile ducts. Bile is a fluid produced by the liver that helps in the digestion and absorption of fats. When bile flow is obstructed or slowed down, it can lead to the accumulation of bile in the liver or bile ducts, causing inflammation, damage to the liver tissue, and potentially leading to other complications. Biliary stasis can be caused by various factors, such as gallstones, inflammation, infection, or structural abnormalities in the bile ducts.", "fig_caption": "Histopathological finding of the liver", "in_text_mention": [{"tokens": "In liver biopsy interphase activity , confluent necrosis , hepatic duct proliferation , and biliary stasis were detected ( Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Fig. 3:"}
-{"question_id": 68, "image": "29354481_trd-2-trd015-g004.jpg", "pair_id": "29354481_trd-2-trd015-g004", "text": "What is the organ shown in the image?\n", "gpt4_answer": "The organ shown in the image is the liver.", "fig_caption": "Gross appearance of cirrhotic liver in Wilson disease.", "in_text_mention": [{"tokens": "Cirrhosis may be micronodular or macronodular ( Fig. 4 ) , or it may be mixed .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 7, "end_tok": 9}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Fig.4"}
-{"question_id": 69, "image": "29354481_trd-2-trd015-g004.jpg", "pair_id": "29354481_trd-2-trd015-g004", "text": "What is the condition of the liver?\n", "gpt4_answer": "The liver in the image appears to be cirrhotic, which means it has undergone significant scarring and structural changes due to chronic liver damage.", "fig_caption": "Gross appearance of cirrhotic liver in Wilson disease.", "in_text_mention": [{"tokens": "Cirrhosis may be micronodular or macronodular ( Fig. 4 ) , or it may be mixed .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 7, "end_tok": 9}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Fig.4"}
-{"question_id": 70, "image": "29354481_trd-2-trd015-g004.jpg", "pair_id": "29354481_trd-2-trd015-g004", "text": "What disease is associated with this liver condition?\n", "gpt4_answer": "The cirrhotic liver in the image is associated with Wilson disease, a rare genetic disorder that causes excessive copper accumulation in the liver, brain, and other vital organs. This accumulation can lead to liver damage and cirrhosis, as seen in the image.", "fig_caption": "Gross appearance of cirrhotic liver in Wilson disease.", "in_text_mention": [{"tokens": "Cirrhosis may be micronodular or macronodular ( Fig. 4 ) , or it may be mixed .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 7, "end_tok": 9}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Fig.4"}
-{"question_id": 71, "image": "29484042_f0015.jpg", "pair_id": "29484042_f0015", "text": "What is the blue arrow pointing to?\n", "gpt4_answer": "The blue arrow in the image is pointing to the stenotic right main bronchus.", "fig_caption": "Fetal MRI SSFSE sequence shows the stenotic right main bronchus (blue arrow) comes from the sloping left main bronchus (red arrow), the position of pseudo carina is low (white arrow).which form a bridging bronchus in type II. LMB, left main bronchus; MRI, magnetic resonance imaging; SSFSE, single-shot fast spin echo.", "in_text_mention": [{"tokens": "On the coronal view , the carina was obviously lower than the aortic arch , which was located at approximately the sixth or seventh thoracic vertebra ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig.\u00a02"}
-{"question_id": 72, "image": "29484042_f0015.jpg", "pair_id": "29484042_f0015", "text": "What is the red arrow pointing to?\n", "gpt4_answer": "The red arrow in the image is pointing to the sloping left main bronchus.", "fig_caption": "Fetal MRI SSFSE sequence shows the stenotic right main bronchus (blue arrow) comes from the sloping left main bronchus (red arrow), the position of pseudo carina is low (white arrow).which form a bridging bronchus in type II. LMB, left main bronchus; MRI, magnetic resonance imaging; SSFSE, single-shot fast spin echo.", "in_text_mention": [{"tokens": "On the coronal view , the carina was obviously lower than the aortic arch , which was located at approximately the sixth or seventh thoracic vertebra ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig.\u00a02"}
-{"question_id": 73, "image": "29484042_f0015.jpg", "pair_id": "29484042_f0015", "text": "What is the white arrow pointing to?\n", "gpt4_answer": "The white arrow in the image is pointing to the position of the pseudo carina, which is described as being low in this case.", "fig_caption": "Fetal MRI SSFSE sequence shows the stenotic right main bronchus (blue arrow) comes from the sloping left main bronchus (red arrow), the position of pseudo carina is low (white arrow).which form a bridging bronchus in type II. LMB, left main bronchus; MRI, magnetic resonance imaging; SSFSE, single-shot fast spin echo.", "in_text_mention": [{"tokens": "On the coronal view , the carina was obviously lower than the aortic arch , which was located at approximately the sixth or seventh thoracic vertebra ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig.\u00a02"}
-{"question_id": 74, "image": "29563712_F5.jpg", "pair_id": "29563712_F5", "text": "What is the size of the eyeball in the image?\n", "gpt4_answer": "The eyeball in the image measures approximately 2.5 cm \u00d7 2.5 cm \u00d7 1.3 cm.", "fig_caption": "Gross specimen of the eyeball of the size 2.5 cm \u00d7 2.5 cm \u00d7 1.3 cm with a gray-white growth attached on the scleral surface, with no involvement of the posterior half of the eyeball and optic nerve", "in_text_mention": [{"tokens": "Outer surface of the eyeball was gray-white with prominent vascular markings [ Figure 5 ] .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 12, "end_tok": 14}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 75, "image": "29563712_F5.jpg", "pair_id": "29563712_F5", "text": "What is the color of the growth on the scleral surface?\n", "gpt4_answer": "The growth on the scleral surface appears to be gray-white in color.", "fig_caption": "Gross specimen of the eyeball of the size 2.5 cm \u00d7 2.5 cm \u00d7 1.3 cm with a gray-white growth attached on the scleral surface, with no involvement of the posterior half of the eyeball and optic nerve", "in_text_mention": [{"tokens": "Outer surface of the eyeball was gray-white with prominent vascular markings [ Figure 5 ] .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 12, "end_tok": 14}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 76, "image": "29563712_F5.jpg", "pair_id": "29563712_F5", "text": "Is the posterior half of the eyeball and optic nerve involved?\n", "gpt4_answer": "According to the information provided, there is no involvement of the posterior half of the eyeball and the optic nerve.", "fig_caption": "Gross specimen of the eyeball of the size 2.5 cm \u00d7 2.5 cm \u00d7 1.3 cm with a gray-white growth attached on the scleral surface, with no involvement of the posterior half of the eyeball and optic nerve", "in_text_mention": [{"tokens": "Outer surface of the eyeball was gray-white with prominent vascular markings [ Figure 5 ] .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 12, "end_tok": 14}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 77, "image": "29774196_F4.jpg", "pair_id": "29774196_F4", "text": "What artery is shown in the image?\n", "gpt4_answer": "The image shows the left middle cerebral artery (MCA).", "fig_caption": "Computed tomography angiography showing almost normal vascularity of the left middle cerebral artery.", "in_text_mention": [{"tokens": "However , the left MCA showed almost normal vascularity on CT angiography ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 13, "end_tok": 15}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "FIGURE 4"}
-{"question_id": 78, "image": "29774196_F4.jpg", "pair_id": "29774196_F4", "text": "What does the image reveal about the vascularity of the artery?\n", "gpt4_answer": "The computed tomography angiography (CTA) image reveals that the vascularity of the left middle cerebral artery appears to be almost normal.", "fig_caption": "Computed tomography angiography showing almost normal vascularity of the left middle cerebral artery.", "in_text_mention": [{"tokens": "However , the left MCA showed almost normal vascularity on CT angiography ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 13, "end_tok": 15}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "FIGURE 4"}
-{"question_id": 79, "image": "29774196_F4.jpg", "pair_id": "29774196_F4", "text": "What is the significance of the normal vascularity?\n", "gpt4_answer": "The normal vascularity of the left middle cerebral artery suggests that there is adequate blood flow through the artery, which is important for supplying oxygen and nutrients to the brain tissue it serves. This finding indicates that there may not be any significant blockages, narrowing, or other abnormalities in the artery that could potentially compromise blood flow and lead to neurological issues or stroke. However, it is important to consider the patient's overall clinical context and consult a healthcare professional for a comprehensive evaluation of their condition.", "fig_caption": "Computed tomography angiography showing almost normal vascularity of the left middle cerebral artery.", "in_text_mention": [{"tokens": "However , the left MCA showed almost normal vascularity on CT angiography ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 13, "end_tok": 15}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "FIGURE 4"}
-{"question_id": 80, "image": "29805352_F2.jpg", "pair_id": "29805352_F2", "text": "What types of cells are present in the portal tracts?\n", "gpt4_answer": "The portal tracts contain abundant CD3+ T cells and scattered PAX8+ B cells.", "fig_caption": "There were also abundant CD3+ T cells (a; \u00d7200) and scattered PAX8+ B cells (b; \u00d7100) within the portal tracts. H&E shows Reed-Sternberg cells (c; \u00d7600) throughout the portal and periportal areas positive for EBV (by Epstein-Barr encoding region in situ hybridization) (d; \u00d7600), CD15 (e; \u00d7600), and CD30 (f; \u00d7600).", "in_text_mention": [{"tokens": "Immunohistochemical stains confirmed that the portal inflammation was composed primarily of CD3 + T lymphocytes , interspersed CD20 + / PAX-5 + B cells , and focal histiocytes ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 29, "end_tok": 31}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Fig. 2."}
-{"question_id": 81, "image": "29805352_F2.jpg", "pair_id": "29805352_F2", "text": "What type of cells are shown in the H&E image?\n", "gpt4_answer": "The H&E image shows Reed-Sternberg cells, which are characteristic of Hodgkin's lymphoma. These cells are large, often multinucleated, and have a distinctive appearance with prominent nucleoli.", "fig_caption": "There were also abundant CD3+ T cells (a; \u00d7200) and scattered PAX8+ B cells (b; \u00d7100) within the portal tracts. H&E shows Reed-Sternberg cells (c; \u00d7600) throughout the portal and periportal areas positive for EBV (by Epstein-Barr encoding region in situ hybridization) (d; \u00d7600), CD15 (e; \u00d7600), and CD30 (f; \u00d7600).", "in_text_mention": [{"tokens": "Immunohistochemical stains confirmed that the portal inflammation was composed primarily of CD3 + T lymphocytes , interspersed CD20 + / PAX-5 + B cells , and focal histiocytes ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 29, "end_tok": 31}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Fig. 2."}
-{"question_id": 82, "image": "29805352_F2.jpg", "pair_id": "29805352_F2", "text": "What markers are the Reed-Sternberg cells positive for?\n", "gpt4_answer": "The Reed-Sternberg cells are positive for EBV (Epstein-Barr virus), CD15, and CD30. These markers are often used to help identify and diagnose Hodgkin's lymphoma.", "fig_caption": "There were also abundant CD3+ T cells (a; \u00d7200) and scattered PAX8+ B cells (b; \u00d7100) within the portal tracts. H&E shows Reed-Sternberg cells (c; \u00d7600) throughout the portal and periportal areas positive for EBV (by Epstein-Barr encoding region in situ hybridization) (d; \u00d7600), CD15 (e; \u00d7600), and CD30 (f; \u00d7600).", "in_text_mention": [{"tokens": "Immunohistochemical stains confirmed that the portal inflammation was composed primarily of CD3 + T lymphocytes , interspersed CD20 + / PAX-5 + B cells , and focal histiocytes ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 29, "end_tok": 31}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Fig. 2."}
-{"question_id": 83, "image": "30210495_F5.jpg", "pair_id": "30210495_F5", "text": "What do the Doppler images show?\n", "gpt4_answer": "The Doppler images show blood flow in the femoral arteries of both wild-type (WT) and BLTR1-deficient (KO) mice at 4 weeks after wire injury (WI). In these color-coded images, red hue indicates regions of maximum perfusion, medium perfusion values are shown in yellow, and lowest perfusion values are represented as blue. Arrows indicate blood flow in an injured femoral artery. The images are representative of 5-6 independent experiments.", "fig_caption": "Involvement of BLTR1 in macrophage infiltration and neointima formation in wire-injured femoral arteries. Doppler images: Blood flow in the femoral arteries of WT and BLTR1-deficient (KO) mice at 4 wks after wire injury (WI) was assessed using a LDPI analyzer. In these color-coded images, red hue indicates regions of maximum perfusion, medium perfusion values are shown in yellow, and lowest perfusion values are represented as blue. Arrows indicate blood flow in an injured femoral artery. Photographs are representative of 5-6 independent experiments. H&E: Cross sections of mouse femoral arteries were prepared at 4 wks after WI, and stained with H&E. HMGB1 and CD36: HMGB1 and macrophage infiltration in the indicated neointima were stained with anti-HMGB1 antibody and anti-CD36 antibody, respectively. \u03b1-SMA: VSMCs were stained with anti-\u03b1-SMA antibody. Images are representative of 5\u20136 independent experiments. Bottom: LDPI ratio was quantified as the ratio of the blue-to-red pixels in the injured artery (WI) vs. non-injured arteries (Control). Neointima volumes in the cross sections of injured femoral artery were determined using an image analyzer. Numbers of HMGB1-positive and CD36-positive cells in neointima area were quantified, and data were presented as the means \u00b1 SEMs of 3-4 independent experiments. **P < 0.01 vs. WT mice.", "in_text_mention": [{"tokens": "Likewise , macrophage infiltration into neointima was also markedly attenuated in BLTR1 deficient mice ( Figure 5 ) , suggesting BLTR1 contributed to vascular inflammation and subsequent neointima formation induced by damage associated mediators secreted in the injured vasculatures .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 15, "end_tok": 17}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 84, "image": "30210495_F5.jpg", "pair_id": "30210495_F5", "text": "What is the purpose of the H&E staining?\n", "gpt4_answer": "The purpose of the H&E (hematoxylin and eosin) staining is to visualize the cross sections of mouse femoral arteries at 4 weeks after wire injury. H&E staining is a common technique used in histology to provide contrast between different tissue components, making it easier to identify and analyze the structure and organization of the tissue. In this case, it helps to assess the neointima formation in the injured femoral arteries.", "fig_caption": "Involvement of BLTR1 in macrophage infiltration and neointima formation in wire-injured femoral arteries. Doppler images: Blood flow in the femoral arteries of WT and BLTR1-deficient (KO) mice at 4 wks after wire injury (WI) was assessed using a LDPI analyzer. In these color-coded images, red hue indicates regions of maximum perfusion, medium perfusion values are shown in yellow, and lowest perfusion values are represented as blue. Arrows indicate blood flow in an injured femoral artery. Photographs are representative of 5-6 independent experiments. H&E: Cross sections of mouse femoral arteries were prepared at 4 wks after WI, and stained with H&E. HMGB1 and CD36: HMGB1 and macrophage infiltration in the indicated neointima were stained with anti-HMGB1 antibody and anti-CD36 antibody, respectively. \u03b1-SMA: VSMCs were stained with anti-\u03b1-SMA antibody. Images are representative of 5\u20136 independent experiments. Bottom: LDPI ratio was quantified as the ratio of the blue-to-red pixels in the injured artery (WI) vs. non-injured arteries (Control). Neointima volumes in the cross sections of injured femoral artery were determined using an image analyzer. Numbers of HMGB1-positive and CD36-positive cells in neointima area were quantified, and data were presented as the means \u00b1 SEMs of 3-4 independent experiments. **P < 0.01 vs. WT mice.", "in_text_mention": [{"tokens": "Likewise , macrophage infiltration into neointima was also markedly attenuated in BLTR1 deficient mice ( Figure 5 ) , suggesting BLTR1 contributed to vascular inflammation and subsequent neointima formation induced by damage associated mediators secreted in the injured vasculatures .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 15, "end_tok": 17}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 85, "image": "30210495_F5.jpg", "pair_id": "30210495_F5", "text": "What do the anti-HMGB1 and anti-CD36 antibodies indicate?\n", "gpt4_answer": "The anti-HMGB1 and anti-CD36 antibodies are used to stain and visualize specific components within the neointima of the injured femoral arteries. The anti-HMGB1 antibody is used to stain and detect the presence of HMGB1, a protein that plays a role in inflammation and tissue damage. The anti-CD36 antibody is used to stain and identify macrophages, which are immune cells involved in inflammation and tissue repair. By staining with these antibodies, the researchers can assess the extent of inflammation and macrophage infiltration in the neointima area, which can provide insights into the role of BLTR1 in vascular inflammation and neointima formation following injury.", "fig_caption": "Involvement of BLTR1 in macrophage infiltration and neointima formation in wire-injured femoral arteries. Doppler images: Blood flow in the femoral arteries of WT and BLTR1-deficient (KO) mice at 4 wks after wire injury (WI) was assessed using a LDPI analyzer. In these color-coded images, red hue indicates regions of maximum perfusion, medium perfusion values are shown in yellow, and lowest perfusion values are represented as blue. Arrows indicate blood flow in an injured femoral artery. Photographs are representative of 5-6 independent experiments. H&E: Cross sections of mouse femoral arteries were prepared at 4 wks after WI, and stained with H&E. HMGB1 and CD36: HMGB1 and macrophage infiltration in the indicated neointima were stained with anti-HMGB1 antibody and anti-CD36 antibody, respectively. \u03b1-SMA: VSMCs were stained with anti-\u03b1-SMA antibody. Images are representative of 5\u20136 independent experiments. Bottom: LDPI ratio was quantified as the ratio of the blue-to-red pixels in the injured artery (WI) vs. non-injured arteries (Control). Neointima volumes in the cross sections of injured femoral artery were determined using an image analyzer. Numbers of HMGB1-positive and CD36-positive cells in neointima area were quantified, and data were presented as the means \u00b1 SEMs of 3-4 independent experiments. **P < 0.01 vs. WT mice.", "in_text_mention": [{"tokens": "Likewise , macrophage infiltration into neointima was also markedly attenuated in BLTR1 deficient mice ( Figure 5 ) , suggesting BLTR1 contributed to vascular inflammation and subsequent neointima formation induced by damage associated mediators secreted in the injured vasculatures .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 15, "end_tok": 17}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 86, "image": "30214911_F4.jpg", "pair_id": "30214911_F4", "text": "What is the size of the mass?\n", "gpt4_answer": "The mass measures 9 cm in its greatest dimension.", "fig_caption": "Gross pathology showing an irregular ovoid mass measuring 9 cm in its greatest dimension, with a smooth serosal surface, prominent vasculature, and a 2.5 \u00d7 1.5 cm central ulcer (arrow).", "in_text_mention": [{"tokens": "Gross pathology revealed a tan-pink irregular ovoid mass measuring 9 cm in its greatest dimension , with a smooth serosal surface , prominent vasculature , and a 2.5 \u00d7 1.5 cm central ulcer ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 34, "end_tok": 36}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 87, "image": "30214911_F4.jpg", "pair_id": "30214911_F4", "text": "What does the arrow point to?\n", "gpt4_answer": "The arrow points to a central ulcer, which is an area of tissue breakdown or erosion on the surface of the mass.", "fig_caption": "Gross pathology showing an irregular ovoid mass measuring 9 cm in its greatest dimension, with a smooth serosal surface, prominent vasculature, and a 2.5 \u00d7 1.5 cm central ulcer (arrow).", "in_text_mention": [{"tokens": "Gross pathology revealed a tan-pink irregular ovoid mass measuring 9 cm in its greatest dimension , with a smooth serosal surface , prominent vasculature , and a 2.5 \u00d7 1.5 cm central ulcer ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 34, "end_tok": 36}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 88, "image": "30214911_F4.jpg", "pair_id": "30214911_F4", "text": "How does the serosal surface appear?\n", "gpt4_answer": "The serosal surface of the mass appears smooth.", "fig_caption": "Gross pathology showing an irregular ovoid mass measuring 9 cm in its greatest dimension, with a smooth serosal surface, prominent vasculature, and a 2.5 \u00d7 1.5 cm central ulcer (arrow).", "in_text_mention": [{"tokens": "Gross pathology revealed a tan-pink irregular ovoid mass measuring 9 cm in its greatest dimension , with a smooth serosal surface , prominent vasculature , and a 2.5 \u00d7 1.5 cm central ulcer ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 34, "end_tok": 36}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 89, "image": "30279704_F3.jpg", "pair_id": "30279704_F3", "text": "What do the black arrows indicate in the left image?\n", "gpt4_answer": "In the left axial T2W MR image of the brain, the black arrows indicate the mild prominence of both lateral ventricles.", "fig_caption": "\nBESS:\n(Left ): Axial T2W MR image of the brain reveals mild prominence of both the lateral ventricles (black arrows) with increased anterior inter-hemispheric distance (green arrow).(Right): Axial T2W MR image of the same infant shows enlarged subarachnoid space along the anterior aspect of the brain (black arrows) with prominent of bilateral lateral ventricles (green arrows). Moreover, the anterior cranio-cortical distance (black arrows) is obviously increased.", "in_text_mention": [{"tokens": "CT and MRI without contrast are also important for evaluating the most common complications associated with external hydrocephalus ( 7 , 38 , 39 ) ( Figure 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 26, "end_tok": 28}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 90, "image": "30279704_F3.jpg", "pair_id": "30279704_F3", "text": "What does the green arrow point to in the left image?\n", "gpt4_answer": "In the left axial T2W MR image, the green arrow points to the increased anterior inter-hemispheric distance.", "fig_caption": "\nBESS:\n(Left ): Axial T2W MR image of the brain reveals mild prominence of both the lateral ventricles (black arrows) with increased anterior inter-hemispheric distance (green arrow).(Right): Axial T2W MR image of the same infant shows enlarged subarachnoid space along the anterior aspect of the brain (black arrows) with prominent of bilateral lateral ventricles (green arrows). Moreover, the anterior cranio-cortical distance (black arrows) is obviously increased.", "in_text_mention": [{"tokens": "CT and MRI without contrast are also important for evaluating the most common complications associated with external hydrocephalus ( 7 , 38 , 39 ) ( Figure 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 26, "end_tok": 28}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 91, "image": "30279704_F3.jpg", "pair_id": "30279704_F3", "text": "What do the black arrows indicate in the right image?\n", "gpt4_answer": "In the right axial T2W MR image of the brain, the black arrows point to the enlarged subarachnoid space along the anterior aspect of the brain, and the increased anterior cranio-cortical distance.", "fig_caption": "\nBESS:\n(Left ): Axial T2W MR image of the brain reveals mild prominence of both the lateral ventricles (black arrows) with increased anterior inter-hemispheric distance (green arrow).(Right): Axial T2W MR image of the same infant shows enlarged subarachnoid space along the anterior aspect of the brain (black arrows) with prominent of bilateral lateral ventricles (green arrows). Moreover, the anterior cranio-cortical distance (black arrows) is obviously increased.", "in_text_mention": [{"tokens": "CT and MRI without contrast are also important for evaluating the most common complications associated with external hydrocephalus ( 7 , 38 , 39 ) ( Figure 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 26, "end_tok": 28}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 92, "image": "30420960_fig2.jpg", "pair_id": "30420960_fig2", "text": "What type of imaging is shown on the left side?\n", "gpt4_answer": "The left side of the image shows an ultrasound image.", "fig_caption": "Ultrasound image (left side) with the corresponding multiplanar reconstructed MR image (right side) of a 55-year-old woman who underwent a previous surgery of the left breast for invasive ductal carcinoma (IDC, pT2N0). After 6 months, a follow-up MR was performed showing a rounded enhancing lesion in the left internal mammary chain (green cross). Second-look ultrasound with coregistration revealed a pathological lymph node.", "in_text_mention": null, "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 93, "image": "30420960_fig2.jpg", "pair_id": "30420960_fig2", "text": "What type of imaging is shown on the right side?\n", "gpt4_answer": "The right side of the image shows a multiplanar reconstructed magnetic resonance (MR) image.", "fig_caption": "Ultrasound image (left side) with the corresponding multiplanar reconstructed MR image (right side) of a 55-year-old woman who underwent a previous surgery of the left breast for invasive ductal carcinoma (IDC, pT2N0). After 6 months, a follow-up MR was performed showing a rounded enhancing lesion in the left internal mammary chain (green cross). Second-look ultrasound with coregistration revealed a pathological lymph node.", "in_text_mention": null, "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 94, "image": "30420960_fig2.jpg", "pair_id": "30420960_fig2", "text": "What is the green cross indicating?\n", "gpt4_answer": "The green cross in the image is indicating a rounded enhancing lesion in the left internal mammary chain. This lesion was identified during a follow-up MR examination after the patient had undergone surgery for invasive ductal carcinoma in the left breast. The second-look ultrasound with coregistration revealed a pathological lymph node, which may be related to the patient's previous breast cancer diagnosis.", "fig_caption": "Ultrasound image (left side) with the corresponding multiplanar reconstructed MR image (right side) of a 55-year-old woman who underwent a previous surgery of the left breast for invasive ductal carcinoma (IDC, pT2N0). After 6 months, a follow-up MR was performed showing a rounded enhancing lesion in the left internal mammary chain (green cross). Second-look ultrasound with coregistration revealed a pathological lymph node.", "in_text_mention": null, "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 95, "image": "30510823_fig1.jpg", "pair_id": "30510823_fig1", "text": "What does the chest X-ray show?\n", "gpt4_answer": "The chest X-ray shows bilateral extensive pneumonic consolidation, which means there is a widespread presence of dense, solidified lung tissue in both lungs. This is often a result of inflammation and accumulation of fluid in the airspaces within the lungs.", "fig_caption": "Chest X-ray of patient with SHS showing bilateral extensive pneumonic consolidation.", "in_text_mention": [{"tokens": "Chest X-ray was suggestive of bilateral , extensive , and patchy consolidation suggestive of infective etiology ( Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 17, "end_tok": 19}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 96, "image": "30510823_fig1.jpg", "pair_id": "30510823_fig1", "text": "What could be the cause of this consolidation?\n", "gpt4_answer": "The consolidation in the chest X-ray is suggestive of an infective etiology, which means it is likely caused by an infection. This could be due to bacterial, viral, or fungal pathogens that have led to inflammation and fluid accumulation in the lungs. Further diagnostic tests and clinical evaluation would be necessary to determine the specific cause and appropriate treatment for the patient.", "fig_caption": "Chest X-ray of patient with SHS showing bilateral extensive pneumonic consolidation.", "in_text_mention": [{"tokens": "Chest X-ray was suggestive of bilateral , extensive , and patchy consolidation suggestive of infective etiology ( Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 17, "end_tok": 19}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 97, "image": "30546931_F9.jpg", "pair_id": "30546931_F9", "text": "What is the age of the patient?\n", "gpt4_answer": "The patient is a 14-year-old female.", "fig_caption": "Viral Sialadenitis. Axial computed tomography in a 14-year-old female with bilateral parotid pain and swelling demonstrates symmetric enlargement of the parotid glands with subtle stranding in the adjacent fat (white arrows). Blood work was positive for parainfluenza virus.", "in_text_mention": [{"tokens": "On imaging , classic findings include bilateral enlargement of the salivary glands , fat stranding , and thickening of the superficial cervical fascia and platysma muscles [ Figure 9 ] . [ 1213 ] Although bilateral involvement is seen in up to 75 % of patients , a small minority of patients may present with only unilateral involvement .", "entities": [{"type": "figure", "entity_id": "9", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 9"}
-{"question_id": 98, "image": "30546931_F9.jpg", "pair_id": "30546931_F9", "text": "What are the white arrows pointing to?\n", "gpt4_answer": "The white arrows in the axial computed tomography (CT) image are pointing to the parotid glands, which appear to be symmetrically enlarged, and the subtle stranding in the adjacent fat.", "fig_caption": "Viral Sialadenitis. Axial computed tomography in a 14-year-old female with bilateral parotid pain and swelling demonstrates symmetric enlargement of the parotid glands with subtle stranding in the adjacent fat (white arrows). Blood work was positive for parainfluenza virus.", "in_text_mention": [{"tokens": "On imaging , classic findings include bilateral enlargement of the salivary glands , fat stranding , and thickening of the superficial cervical fascia and platysma muscles [ Figure 9 ] . [ 1213 ] Although bilateral involvement is seen in up to 75 % of patients , a small minority of patients may present with only unilateral involvement .", "entities": [{"type": "figure", "entity_id": "9", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 9"}
-{"question_id": 99, "image": "30546931_F9.jpg", "pair_id": "30546931_F9", "text": "What is the diagnosis?\n", "gpt4_answer": "The diagnosis for this patient is viral sialadenitis, which is an inflammation of the salivary glands caused by a viral infection. In this case, the blood work was positive for the parainfluenza virus. The CT image shows classic findings of bilateral enlargement of the salivary glands, fat stranding, and thickening of the superficial cervical fascia and platysma muscles.", "fig_caption": "Viral Sialadenitis. Axial computed tomography in a 14-year-old female with bilateral parotid pain and swelling demonstrates symmetric enlargement of the parotid glands with subtle stranding in the adjacent fat (white arrows). Blood work was positive for parainfluenza virus.", "in_text_mention": [{"tokens": "On imaging , classic findings include bilateral enlargement of the salivary glands , fat stranding , and thickening of the superficial cervical fascia and platysma muscles [ Figure 9 ] . [ 1213 ] Although bilateral involvement is seen in up to 75 % of patients , a small minority of patients may present with only unilateral involvement .", "entities": [{"type": "figure", "entity_id": "9", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 9"}
-{"question_id": 100, "image": "30910324_fig3.jpg", "pair_id": "30910324_fig3", "text": "What is the main focus of the MRI image?\n", "gpt4_answer": "The main focus of the MRI image is a fetus with a double aortic arch at 32 weeks of gestation. The image shows the descending aorta (DAo), arterial duct (D), and left (L) and right (R) aortic arches.", "fig_caption": "Motion-corrected MRI data from a fetus with double aortic arch at 32 weeks Shown are the descending aorta (DAo), arterial duct (D), and left (L) and right (R) aortic arches. At 2 months postnatal age, contrast-enhanced MRI could show a right-sided arch (middle panel); however, a ligamentous remnant of the left arch was predicted on the basis of the fetal MRI findings (asterisk); this finding was confirmed at surgery (right panel). The distal remnant of the arterial duct\u2014analogous to the diverticulum of Kommerell\u2014is also seen (K). See video 3 for more detail.", "in_text_mention": [{"tokens": "Example images from 3D reconstructed data are shown in Figure 3 , Figure 4 , Figure 5 ; appendix p 3 ; videos 2-4 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 9, "end_tok": 11}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 101, "image": "30910324_fig3.jpg", "pair_id": "30910324_fig3", "text": "What is the significance of the asterisk?\n", "gpt4_answer": "The asterisk in the image is used to indicate a predicted ligamentous remnant of the left aortic arch. This prediction was based on the fetal MRI findings, and it was later confirmed during surgery.", "fig_caption": "Motion-corrected MRI data from a fetus with double aortic arch at 32 weeks Shown are the descending aorta (DAo), arterial duct (D), and left (L) and right (R) aortic arches. At 2 months postnatal age, contrast-enhanced MRI could show a right-sided arch (middle panel); however, a ligamentous remnant of the left arch was predicted on the basis of the fetal MRI findings (asterisk); this finding was confirmed at surgery (right panel). The distal remnant of the arterial duct\u2014analogous to the diverticulum of Kommerell\u2014is also seen (K). See video 3 for more detail.", "in_text_mention": [{"tokens": "Example images from 3D reconstructed data are shown in Figure 3 , Figure 4 , Figure 5 ; appendix p 3 ; videos 2-4 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 9, "end_tok": 11}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 102, "image": "30910324_fig3.jpg", "pair_id": "30910324_fig3", "text": "What is the diverticulum of Kommerell?\n", "gpt4_answer": "The diverticulum of Kommerell is a rare anatomical variant in which an aberrant blood vessel arises from the aortic arch. In the context of the image, the distal remnant of the arterial duct is analogous to the diverticulum of Kommerell. This means that the structure seen in the image is similar to the diverticulum of Kommerell, which is a vascular anomaly that can be associated with a double aortic arch.", "fig_caption": "Motion-corrected MRI data from a fetus with double aortic arch at 32 weeks Shown are the descending aorta (DAo), arterial duct (D), and left (L) and right (R) aortic arches. At 2 months postnatal age, contrast-enhanced MRI could show a right-sided arch (middle panel); however, a ligamentous remnant of the left arch was predicted on the basis of the fetal MRI findings (asterisk); this finding was confirmed at surgery (right panel). The distal remnant of the arterial duct\u2014analogous to the diverticulum of Kommerell\u2014is also seen (K). See video 3 for more detail.", "in_text_mention": [{"tokens": "Example images from 3D reconstructed data are shown in Figure 3 , Figure 4 , Figure 5 ; appendix p 3 ; videos 2-4 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 9, "end_tok": 11}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 103, "image": "30911703_fig1.jpg", "pair_id": "30911703_fig1", "text": "What type of aneurysm is shown in the image?\n", "gpt4_answer": "The image shows a saccular aneurysm of the right peroneal artery.", "fig_caption": "Computed tomography scan of the right peroneal artery aneurysm.", "in_text_mention": [{"tokens": "A computed tomography scan with contrast of the right lower extremity demonstrated a 2.7- \u00d7 3.2- \u00d7 3.2-cm saccular aneurysm of his right peroneal artery just distal to the tibioperoneal trunk ( Fig 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 32, "end_tok": 34}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig\u00a01"}
-{"question_id": 104, "image": "30911703_fig1.jpg", "pair_id": "30911703_fig1", "text": "Where is the aneurysm located?\n", "gpt4_answer": "The aneurysm is located just distal to the tibioperoneal trunk in the right peroneal artery.", "fig_caption": "Computed tomography scan of the right peroneal artery aneurysm.", "in_text_mention": [{"tokens": "A computed tomography scan with contrast of the right lower extremity demonstrated a 2.7- \u00d7 3.2- \u00d7 3.2-cm saccular aneurysm of his right peroneal artery just distal to the tibioperoneal trunk ( Fig 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 32, "end_tok": 34}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig\u00a01"}
-{"question_id": 105, "image": "30911703_fig1.jpg", "pair_id": "30911703_fig1", "text": "What imaging technique was used?\n", "gpt4_answer": "A computed tomography (CT) scan with contrast was used to visualize the aneurysm in the right peroneal artery.", "fig_caption": "Computed tomography scan of the right peroneal artery aneurysm.", "in_text_mention": [{"tokens": "A computed tomography scan with contrast of the right lower extremity demonstrated a 2.7- \u00d7 3.2- \u00d7 3.2-cm saccular aneurysm of his right peroneal artery just distal to the tibioperoneal trunk ( Fig 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 32, "end_tok": 34}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig\u00a01"}
-{"question_id": 106, "image": "30984437_fig2.jpg", "pair_id": "30984437_fig2", "text": "What does the chest CT show?\n", "gpt4_answer": "The chest CT image shows bilateral airspace consolidation and ground-glass opacity. These findings suggest that there are areas of increased density in the lung tissue on both sides, which could be due to various reasons such as infection, inflammation, or other lung conditions.", "fig_caption": "Chest computed tomography revealed bilateral airspace consolidation and ground-glass opacity.", "in_text_mention": [{"tokens": "Chest computed tomography ( CT ) revealed bilateral airspace consolidation and ground-glass opacity ( Figure 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 14, "end_tok": 16}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 107, "image": "30984437_fig2.jpg", "pair_id": "30984437_fig2", "text": "What is ground-glass opacity?\n", "gpt4_answer": "Ground-glass opacity is a term used to describe a specific appearance on a chest CT scan. It refers to an area of hazy increased lung opacity, which means that the lung tissue appears less transparent than normal. This appearance is usually caused by a partial filling of airspaces, thickening of the walls of the airspaces, or a combination of both. Ground-glass opacities can be associated with various lung conditions, including infections, inflammation, interstitial lung diseases, and even some types of lung cancer.", "fig_caption": "Chest computed tomography revealed bilateral airspace consolidation and ground-glass opacity.", "in_text_mention": [{"tokens": "Chest computed tomography ( CT ) revealed bilateral airspace consolidation and ground-glass opacity ( Figure 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 14, "end_tok": 16}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 108, "image": "30984437_fig2.jpg", "pair_id": "30984437_fig2", "text": "What could be the possible causes of bilateral airspace consolidation?\n", "gpt4_answer": "Bilateral airspace consolidation can be caused by a variety of factors. Some possible causes include:\n\n1. Infections: Bacterial, viral, or fungal infections can lead to inflammation and fluid accumulation in the lung tissue, resulting in consolidation.\n2. Pneumonia: This is a common cause of consolidation, as the infection leads to inflammation and fluid build-up in the alveoli (air sacs) of the lungs.\n3. Pulmonary edema: Fluid accumulation in the lungs due to heart failure, kidney disease, or other conditions can cause bilateral airspace consolidation.\n4. Acute respiratory distress syndrome (ARDS): This is a severe lung condition that can result from various causes, such as sepsis, trauma, or severe pneumonia, and leads to widespread inflammation and consolidation in the lungs.\n5. Interstitial lung diseases: These are a group of lung disorders that cause inflammation and scarring of the lung tissue, which can lead to consolidation.\n\nIt is important to consider the patient's clinical history, symptoms, and other diagnostic tests to determine the specific cause of bilateral airspace consolidation. A healthcare professional should be consulted for a thorough evaluation and proper diagnosis.", "fig_caption": "Chest computed tomography revealed bilateral airspace consolidation and ground-glass opacity.", "in_text_mention": [{"tokens": "Chest computed tomography ( CT ) revealed bilateral airspace consolidation and ground-glass opacity ( Figure 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 14, "end_tok": 16}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 2"}
-{"question_id": 109, "image": "31404134_F5.jpg", "pair_id": "31404134_F5", "text": "Is there any pleural effusion visible in the image?\n", "gpt4_answer": "No, the chest x-ray image shows no pleural effusion.", "fig_caption": "Chest x-ray, few weeks after starting hemodialysis, showed no pleural effusion. Perm catheter is in place", "in_text_mention": [{"tokens": "Follow-up evaluation showed neither recurrence of symptoms nor recurrence of pleural effusion on chest x-ray ( see Figure 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 17, "end_tok": 19}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 110, "image": "31404134_F5.jpg", "pair_id": "31404134_F5", "text": "What medical device is in place?\n", "gpt4_answer": "A perm catheter is in place, as indicated in the image.", "fig_caption": "Chest x-ray, few weeks after starting hemodialysis, showed no pleural effusion. Perm catheter is in place", "in_text_mention": [{"tokens": "Follow-up evaluation showed neither recurrence of symptoms nor recurrence of pleural effusion on chest x-ray ( see Figure 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 17, "end_tok": 19}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 111, "image": "31404134_F5.jpg", "pair_id": "31404134_F5", "text": "What is the purpose of the perm catheter?\n", "gpt4_answer": "A perm catheter, also known as a permcath or a tunneled dialysis catheter, is a type of long-term catheter used for hemodialysis. It is inserted into a large vein, typically in the chest or neck, and is tunneled under the skin to create a stable and secure access point for hemodialysis treatments. This allows for the efficient removal of waste products and excess fluid from the blood during dialysis sessions, which is essential for patients with kidney failure or other conditions that impair kidney function.", "fig_caption": "Chest x-ray, few weeks after starting hemodialysis, showed no pleural effusion. Perm catheter is in place", "in_text_mention": [{"tokens": "Follow-up evaluation showed neither recurrence of symptoms nor recurrence of pleural effusion on chest x-ray ( see Figure 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 17, "end_tok": 19}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 112, "image": "32009718_F1.jpg", "pair_id": "32009718_F1", "text": "What is the main finding in the chest X-ray?\n", "gpt4_answer": "The main finding in the chest X-ray is the presence of patchy areas of consolidation throughout the right lung and in the left upper zone.", "fig_caption": "Chest X-ray of a patient at presentation. Patchy areas of consolidation seen throughout the right lung and left upper zone", "in_text_mention": [{"tokens": "Chest roentgenogram showed patchy areas of consolidation throughout the right lung fields and left upper lobe [ Figure 1 ] .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 17, "end_tok": 19}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 113, "image": "32009718_F1.jpg", "pair_id": "32009718_F1", "text": "What does consolidation mean in this context?\n", "gpt4_answer": "In the context of a chest X-ray, consolidation refers to an area of the lung where the air spaces (alveoli) are filled with fluid, pus, blood, or cells, making the lung tissue appear more solid and dense. This can be caused by various conditions, such as pneumonia, pulmonary edema, or lung injury. Consolidation can be seen as patchy or diffuse areas of increased opacity on the X-ray image.", "fig_caption": "Chest X-ray of a patient at presentation. Patchy areas of consolidation seen throughout the right lung and left upper zone", "in_text_mention": [{"tokens": "Chest roentgenogram showed patchy areas of consolidation throughout the right lung fields and left upper lobe [ Figure 1 ] .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 17, "end_tok": 19}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 114, "image": "32009718_F1.jpg", "pair_id": "32009718_F1", "text": "What could be the possible cause of these consolidations?\n", "gpt4_answer": "There are several possible causes for the consolidations seen in the chest X-ray. Some common causes include pneumonia (infection of the lung tissue), pulmonary edema (fluid accumulation in the lungs), or lung injury due to trauma or other factors. It is important to consider the patient's clinical history, symptoms, and other diagnostic tests to determine the exact cause of the consolidations and provide appropriate treatment. A healthcare professional should be consulted for a thorough evaluation and proper diagnosis.", "fig_caption": "Chest X-ray of a patient at presentation. Patchy areas of consolidation seen throughout the right lung and left upper zone", "in_text_mention": [{"tokens": "Chest roentgenogram showed patchy areas of consolidation throughout the right lung fields and left upper lobe [ Figure 1 ] .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 17, "end_tok": 19}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 115, "image": "32418875_fig1.jpg", "pair_id": "32418875_fig1", "text": "What is the purpose of the pathway shown in the image?\n", "gpt4_answer": "The purpose of the pathway shown in the image is to provide a systematic approach for screening adult COVID-19 patients admitted to the hospital for the risk of acute heart failure and circulatory collapse. This pathway aims to balance the need for identifying patients at risk while also protecting healthcare workers from virus exposure and preserving personal protective equipment (PPE).", "fig_caption": "Adult pathway to screen for acute heart failure in COVID-19 patients admitted to hospital. Abbreviations: ED, emergency department; EUC, electrolytes, urea and creatine; LFT, liver function test; CRP, C-reactive protein; LDH, lactate dehydrogenase; CXR, chest X-ray; ECG, electrocardiograph; HDU, high dependency unit; ICU, intensive care unit; TTE, transthoracic echocardiogram; LV, left ventricular.", "in_text_mention": [{"tokens": "We suggest the following algorithm ( Figure 1 ) , to better identify COVID-19 patients at risk for severe heart failure and circulatory collapse , whilst balancing the need to protect health care workers from virus exposure and to preserve personal protective equipment ( PPE ) :", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 6, "end_tok": 8}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure\u00a01"}
-{"question_id": 116, "image": "32418875_fig1.jpg", "pair_id": "32418875_fig1", "text": "What are some of the tests and procedures included in the pathway?\n", "gpt4_answer": "The pathway includes various tests and procedures such as electrolytes, urea, and creatine (EUC) tests, liver function tests (LFT), C-reactive protein (CRP) tests, lactate dehydrogenase (LDH) tests, chest X-rays (CXR), electrocardiographs (ECG), and transthoracic echocardiograms (TTE). Additionally, it involves different levels of care, such as the emergency department (ED), high dependency unit (HDU), and intensive care unit (ICU).", "fig_caption": "Adult pathway to screen for acute heart failure in COVID-19 patients admitted to hospital. Abbreviations: ED, emergency department; EUC, electrolytes, urea and creatine; LFT, liver function test; CRP, C-reactive protein; LDH, lactate dehydrogenase; CXR, chest X-ray; ECG, electrocardiograph; HDU, high dependency unit; ICU, intensive care unit; TTE, transthoracic echocardiogram; LV, left ventricular.", "in_text_mention": [{"tokens": "We suggest the following algorithm ( Figure 1 ) , to better identify COVID-19 patients at risk for severe heart failure and circulatory collapse , whilst balancing the need to protect health care workers from virus exposure and to preserve personal protective equipment ( PPE ) :", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 6, "end_tok": 8}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure\u00a01"}
-{"question_id": 117, "image": "32418875_fig1.jpg", "pair_id": "32418875_fig1", "text": "What is the main focus of the pathway in relation to COVID-19 patients?\n", "gpt4_answer": "The main focus of the pathway in relation to COVID-19 patients is to identify those who are at risk for severe heart failure and circulatory collapse. By following this pathway, healthcare professionals can better assess the patients' heart function and provide appropriate care while minimizing the risk of virus exposure for healthcare workers and preserving personal protective equipment.", "fig_caption": "Adult pathway to screen for acute heart failure in COVID-19 patients admitted to hospital. Abbreviations: ED, emergency department; EUC, electrolytes, urea and creatine; LFT, liver function test; CRP, C-reactive protein; LDH, lactate dehydrogenase; CXR, chest X-ray; ECG, electrocardiograph; HDU, high dependency unit; ICU, intensive care unit; TTE, transthoracic echocardiogram; LV, left ventricular.", "in_text_mention": [{"tokens": "We suggest the following algorithm ( Figure 1 ) , to better identify COVID-19 patients at risk for severe heart failure and circulatory collapse , whilst balancing the need to protect health care workers from virus exposure and to preserve personal protective equipment ( PPE ) :", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 6, "end_tok": 8}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure\u00a01"}
-{"question_id": 118, "image": "32548036_F5.jpg", "pair_id": "32548036_F5", "text": "Can you describe the appearance of the gross specimen?\n", "gpt4_answer": "I'm sorry, but I cannot provide a detailed description of the gross specimen without more information or access to the image. However, a gross specimen generally refers to a sample of tissue or an organ that has been removed from the body for examination, usually during a surgical procedure or biopsy. The appearance of the specimen can vary greatly depending on the type of tissue, the presence of any abnormalities, and the specific context of the image. If you could provide more information about the specimen or its context, I would be happy to help answer any questions you may have.", "fig_caption": "Gross specimen.", "in_text_mention": null, "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Figure 5"}
-{"question_id": 119, "image": "32715172_fig2.jpg", "pair_id": "32715172_fig2", "text": "What type of imaging technique is used in this figure?\n", "gpt4_answer": "The imaging technique used in this figure is aortography, which is a type of angiography that specifically visualizes the aorta and its branches.", "fig_caption": "Preoperative aortography demonstrating anatomy consistent with the computed tomography scan.", "in_text_mention": [{"tokens": "Intraoperative flush abdominal aortography confirmed computed tomography angiography findings and a suitable anatomy for the proposed repair ( Fig 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 18, "end_tok": 20}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig\u00a02"}
-{"question_id": 120, "image": "32715172_fig2.jpg", "pair_id": "32715172_fig2", "text": "What does the image show?\n", "gpt4_answer": "The image shows the anatomy of the aorta and its branches. It demonstrates that the anatomy is consistent with the findings from a computed tomography (CT) scan. This consistency helps confirm the suitability of the patient's anatomy for the proposed repair.", "fig_caption": "Preoperative aortography demonstrating anatomy consistent with the computed tomography scan.", "in_text_mention": [{"tokens": "Intraoperative flush abdominal aortography confirmed computed tomography angiography findings and a suitable anatomy for the proposed repair ( Fig 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 18, "end_tok": 20}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig\u00a02"}
-{"question_id": 121, "image": "32715172_fig2.jpg", "pair_id": "32715172_fig2", "text": "What was the purpose of this imaging technique?\n", "gpt4_answer": "The purpose of the aortography in this context was to confirm the findings from a computed tomography angiography (CTA) and to ensure that the patient's anatomy was suitable for the proposed repair. By visualizing the aorta and its branches, the medical team can better plan and prepare for the surgical procedure, reducing the risk of complications and improving the overall outcome.", "fig_caption": "Preoperative aortography demonstrating anatomy consistent with the computed tomography scan.", "in_text_mention": [{"tokens": "Intraoperative flush abdominal aortography confirmed computed tomography angiography findings and a suitable anatomy for the proposed repair ( Fig 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 18, "end_tok": 20}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Fig\u00a02"}
-{"question_id": 122, "image": "32867401_materials-13-03795-f015.jpg", "pair_id": "32867401_materials-13-03795-f015", "text": "What is the purpose of the multi-modal Janus nanoparticulate system?\n", "gpt4_answer": "The multi-modal Janus nanoparticulate system is designed to combine different imaging and targeting capabilities within a single nanoparticle. In this case, the system is composed of a SPION/MSN core/shell face and an Au nanoparticle face. The SPION (superparamagnetic iron oxide nanoparticle) component provides T2 MRI contrast capabilities, while the Au (gold) nanoparticles enable CT imaging. Additionally, the system is functionalized with a fluorescent dye (Alexa Fluor\u00ae 647) for optical imaging and a tumor-targeting peptide (cRDG) for active tumor targeting. This combination of features allows for enhanced imaging and targeted delivery of therapeutics to tumors, potentially improving the diagnosis and treatment of cancer.", "fig_caption": "A schematic representing a multi-modal Janus nanoparticulate system composed of a SPION/MSN core/shell face and Au nanoparticle face. T2 MRI contrast capabilities arise from the encapsulated SPION with Au nanoparticles providing the possibility for CT imaging. As shown in this schematic, the system is functionalised with a fluorescent dye (Alexa Fluor\u00ae 647) for optical imaging in addition to cRDG for active tumour targeting. Reprinted with permission from reference [178]. Copyright (2018) American Chemical Society.", "in_text_mention": [{"tokens": "The gold interface was modified with a fluorescent dye ( Alexa Fluor\u00ae 647 ) and the MSN face with a tumour targeting peptide cRDG ( Figure 15 ) .", "entities": [{"type": "figure", "entity_id": "15", "start_tok": 25, "end_tok": 27}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 15"}
-{"question_id": 123, "image": "32952957_F4.jpg", "pair_id": "32952957_F4", "text": "What is the purpose of Sirius Red staining?\n", "gpt4_answer": "Sirius Red staining is a histological technique used to visualize and quantify collagen fibers in tissue samples. Collagen is a major component of the extracellular matrix and plays a crucial role in maintaining the structural integrity of tissues. By staining collagen fibers with Sirius Red, researchers can assess the presence, distribution, and extent of fibrosis (excessive accumulation of collagen) in various tissues, which can be indicative of pathological changes or disease progression.", "fig_caption": "Absence of signs of fibrosis in the left ventricles of 3, 5, 3\u2019 triiodo-L-thyronine treated rats. (top) Representative microphotographs of histological slices stained with Sirius Red of hearts from short-term T3 treated and vehicle-treated rats (red color indicates collagen fibers; scale bar 25 \u00b5m). (bottom) Quantitative analysis of left ventricular collagen-positive areas in each of the different animal groups. The positively stained collagen area was expressed as a percentage of the total area. Short-term hyperthyroidism was induced by the daily subcutaneous injection of T3 (500 \u03bcg.kg-1) for 1 (T31d), 3 (T33d), and 10 (T310d) days. Control animals were injected daily with the vehicle for 1 (V1d), 3 (V3d), and 10 (V10d) days. Values are means\u00b1SEM. P not significant; n= 60 randomly chosen frames from 6 hearts per group (one-way ANOVA)", "in_text_mention": [{"tokens": "Sirius Red staining to determine collagen deposition showed no more connective tissue in the left ventricles of T31d , T33d , and T310d rats compared with their respective temporal controls ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 31, "end_tok": 33}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 124, "image": "32952957_F4.jpg", "pair_id": "32952957_F4", "text": "What is the scale bar in the microphotographs?\n", "gpt4_answer": "The scale bar in the microphotographs is 25 \u00b5m (micrometers). This scale bar provides a reference for the size of the structures in the image, allowing you to estimate the dimensions of the collagen fibers and other features in the tissue sample.", "fig_caption": "Absence of signs of fibrosis in the left ventricles of 3, 5, 3\u2019 triiodo-L-thyronine treated rats. (top) Representative microphotographs of histological slices stained with Sirius Red of hearts from short-term T3 treated and vehicle-treated rats (red color indicates collagen fibers; scale bar 25 \u00b5m). (bottom) Quantitative analysis of left ventricular collagen-positive areas in each of the different animal groups. The positively stained collagen area was expressed as a percentage of the total area. Short-term hyperthyroidism was induced by the daily subcutaneous injection of T3 (500 \u03bcg.kg-1) for 1 (T31d), 3 (T33d), and 10 (T310d) days. Control animals were injected daily with the vehicle for 1 (V1d), 3 (V3d), and 10 (V10d) days. Values are means\u00b1SEM. P not significant; n= 60 randomly chosen frames from 6 hearts per group (one-way ANOVA)", "in_text_mention": [{"tokens": "Sirius Red staining to determine collagen deposition showed no more connective tissue in the left ventricles of T31d , T33d , and T310d rats compared with their respective temporal controls ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 31, "end_tok": 33}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 125, "image": "32952957_F4.jpg", "pair_id": "32952957_F4", "text": "What is the main finding in the left ventricles of the treated rats?\n", "gpt4_answer": "The main finding in the left ventricles of the treated rats is the absence of signs of fibrosis. The Sirius Red staining showed no significant increase in collagen deposition in the left ventricles of the short-term T3 treated rats compared to their respective temporal controls. This suggests that the short-term hyperthyroidism induced by T3 treatment did not lead to an excessive accumulation of collagen fibers in the left ventricles of the rat hearts.", "fig_caption": "Absence of signs of fibrosis in the left ventricles of 3, 5, 3\u2019 triiodo-L-thyronine treated rats. (top) Representative microphotographs of histological slices stained with Sirius Red of hearts from short-term T3 treated and vehicle-treated rats (red color indicates collagen fibers; scale bar 25 \u00b5m). (bottom) Quantitative analysis of left ventricular collagen-positive areas in each of the different animal groups. The positively stained collagen area was expressed as a percentage of the total area. Short-term hyperthyroidism was induced by the daily subcutaneous injection of T3 (500 \u03bcg.kg-1) for 1 (T31d), 3 (T33d), and 10 (T310d) days. Control animals were injected daily with the vehicle for 1 (V1d), 3 (V3d), and 10 (V10d) days. Values are means\u00b1SEM. P not significant; n= 60 randomly chosen frames from 6 hearts per group (one-way ANOVA)", "in_text_mention": [{"tokens": "Sirius Red staining to determine collagen deposition showed no more connective tissue in the left ventricles of T31d , T33d , and T310d rats compared with their respective temporal controls ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 31, "end_tok": 33}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 126, "image": "33708306_f0004.jpg", "pair_id": "33708306_f0004", "text": "What does the white arrow indicate?\n", "gpt4_answer": "The white arrow in the chest X-ray image is pointing to an area of mediastinal widening.", "fig_caption": "Chest X ray - case 2: mediastinal widening (white arrow)", "in_text_mention": [{"tokens": "Chest X ray showed a mediastinal widening ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 8, "end_tok": 10}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 127, "image": "33708306_f0004.jpg", "pair_id": "33708306_f0004", "text": "What does mediastinal widening mean?\n", "gpt4_answer": "Mediastinal widening refers to an abnormal increase in the width of the mediastinum, which is the central compartment of the thoracic cavity. It is located between the lungs and contains vital structures such as the heart, major blood vessels, trachea, and esophagus. Mediastinal widening can be caused by various factors, such as enlarged lymph nodes, tumors, infections, or aneurysms. It is an important finding on a chest X-ray that may indicate an underlying medical condition that requires further investigation and evaluation.", "fig_caption": "Chest X ray - case 2: mediastinal widening (white arrow)", "in_text_mention": [{"tokens": "Chest X ray showed a mediastinal widening ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 8, "end_tok": 10}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 4"}
-{"question_id": 128, "image": "33864678_f1-amjcaserep-22-e930902.jpg", "pair_id": "33864678_f1-amjcaserep-22-e930902", "text": "What is the origin of the mass?\n", "gpt4_answer": "The mass appears to originate from the posterior costal arc of the eighth rib on the patient's left side.", "fig_caption": "A computed tomography scan shows that the mass arises from the posterior costal arc of the eighth rib on the patient\u2019s left side and does not involve the medulla.", "in_text_mention": null, "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1."}
-{"question_id": 129, "image": "33864678_f1-amjcaserep-22-e930902.jpg", "pair_id": "33864678_f1-amjcaserep-22-e930902", "text": "Does the mass involve the medulla?\n", "gpt4_answer": "No, the computed tomography scan shows that the mass does not involve the medulla.", "fig_caption": "A computed tomography scan shows that the mass arises from the posterior costal arc of the eighth rib on the patient\u2019s left side and does not involve the medulla.", "in_text_mention": null, "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1."}
-{"question_id": 130, "image": "33864678_f1-amjcaserep-22-e930902.jpg", "pair_id": "33864678_f1-amjcaserep-22-e930902", "text": "What type of imaging technique is used?\n", "gpt4_answer": "A computed tomography (CT) scan is the imaging technique used in this case. CT scans use X-ray technology to create detailed cross-sectional images of the body, which can help visualize and diagnose various conditions, including the presence of masses or tumors.", "fig_caption": "A computed tomography scan shows that the mass arises from the posterior costal arc of the eighth rib on the patient\u2019s left side and does not involve the medulla.", "in_text_mention": null, "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1."}
-{"question_id": 131, "image": "33935946_F1.jpg", "pair_id": "33935946_F1", "text": "What type of imaging technique is used in this image?\n", "gpt4_answer": "The image is from a multi-modal MRI (Magnetic Resonance Imaging) dataset.", "fig_caption": "Selected slice from a multi-modal MRI dataset of a patient with Parkinson's disease.", "in_text_mention": [{"tokens": "All participants were scanned at the University Medical Center Hamburg-Eppendorf , Germany , using a 3T Siemens Skyra MR scanner ( Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 21, "end_tok": 23}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 132, "image": "33935946_F1.jpg", "pair_id": "33935946_F1", "text": "What is the condition of the patient?\n", "gpt4_answer": "The patient in the image has Parkinson's disease.", "fig_caption": "Selected slice from a multi-modal MRI dataset of a patient with Parkinson's disease.", "in_text_mention": [{"tokens": "All participants were scanned at the University Medical Center Hamburg-Eppendorf , Germany , using a 3T Siemens Skyra MR scanner ( Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 21, "end_tok": 23}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 133, "image": "33935946_F1.jpg", "pair_id": "33935946_F1", "text": "What is the strength of the MRI scanner used?\n", "gpt4_answer": "The MRI scanner used for this image is a 3 Tesla (3T) scanner.", "fig_caption": "Selected slice from a multi-modal MRI dataset of a patient with Parkinson's disease.", "in_text_mention": [{"tokens": "All participants were scanned at the University Medical Center Hamburg-Eppendorf , Germany , using a 3T Siemens Skyra MR scanner ( Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 21, "end_tok": 23}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 1"}
-{"question_id": 134, "image": "34026556_fig2.jpg", "pair_id": "34026556_fig2", "text": "What is the relationship between the appendix and the right kidney in the image?\n", "gpt4_answer": "In the image, the appendix appears to be adhered to the right kidney. This is an abnormal finding, as the appendix and the kidney are typically not directly connected or attached to each other.", "fig_caption": "Gross image of appendix adhered to right kidney.", "in_text_mention": [{"tokens": "An appendectomy was performed to allow for complete removal of the fistulous tract along with the right kidney ( Fig. 2 , Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Fig. 2"}
-{"question_id": 135, "image": "34026556_fig2.jpg", "pair_id": "34026556_fig2", "text": "What procedure was performed?\n", "gpt4_answer": "An appendectomy was performed, which is a surgical procedure to remove the appendix. In this particular case, the appendectomy was done to allow for complete removal of the fistulous tract along with the right kidney.", "fig_caption": "Gross image of appendix adhered to right kidney.", "in_text_mention": [{"tokens": "An appendectomy was performed to allow for complete removal of the fistulous tract along with the right kidney ( Fig. 2 , Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Fig. 2"}
-{"question_id": 136, "image": "34026556_fig2.jpg", "pair_id": "34026556_fig2", "text": "What is a fistulous tract?\n", "gpt4_answer": "A fistulous tract is an abnormal connection or channel between two organs or between an organ and the skin. It can be caused by various factors, such as infection, inflammation, injury, or a disease process. In this specific case, the fistulous tract seems to be related to the abnormal adhesion between the appendix and the right kidney. The surgical removal of the appendix and the right kidney was performed to address this issue.", "fig_caption": "Gross image of appendix adhered to right kidney.", "in_text_mention": [{"tokens": "An appendectomy was performed to allow for complete removal of the fistulous tract along with the right kidney ( Fig. 2 , Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "conversation", "fig_label": "Fig. 2"}
-{"question_id": 137, "image": "34267836_F7.jpg", "pair_id": "34267836_F7", "text": "What type of tissue is shown in the image?\n", "gpt4_answer": "The image shows a metastatic liver lesion, which is a tissue sample from the liver containing cancer cells that have spread from another part of the body.", "fig_caption": "Mixed adenocarcinoma high-grade neuroendocrine carcinoma in metastatic liver lesion (H&E, original \u00d7 400). H&E: hematoxylin and eosin stain.", "in_text_mention": [{"tokens": "Liver biopsy also showed mixed adenocarcinoma and neuroendocrine carcinoma , high-grade ( Fig. 7 ) .", "entities": [{"type": "figure", "entity_id": "7", "start_tok": 12, "end_tok": 14}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 7"}
-{"question_id": 138, "image": "34267836_F7.jpg", "pair_id": "34267836_F7", "text": "What type of carcinoma is present?\n", "gpt4_answer": "The image shows a mixed adenocarcinoma and high-grade neuroendocrine carcinoma in the metastatic liver lesion.", "fig_caption": "Mixed adenocarcinoma high-grade neuroendocrine carcinoma in metastatic liver lesion (H&E, original \u00d7 400). H&E: hematoxylin and eosin stain.", "in_text_mention": [{"tokens": "Liver biopsy also showed mixed adenocarcinoma and neuroendocrine carcinoma , high-grade ( Fig. 7 ) .", "entities": [{"type": "figure", "entity_id": "7", "start_tok": 12, "end_tok": 14}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 7"}
-{"question_id": 139, "image": "34267836_F7.jpg", "pair_id": "34267836_F7", "text": "What type of stain is used?\n", "gpt4_answer": "The stain used in this image is hematoxylin and eosin (H&E). This is a common staining technique used in histology to visualize the structure and features of tissue samples. Hematoxylin stains cell nuclei blue, while eosin stains the cytoplasm and extracellular matrix pink. This contrast helps to differentiate various cell types and structures within the tissue.", "fig_caption": "Mixed adenocarcinoma high-grade neuroendocrine carcinoma in metastatic liver lesion (H&E, original \u00d7 400). H&E: hematoxylin and eosin stain.", "in_text_mention": [{"tokens": "Liver biopsy also showed mixed adenocarcinoma and neuroendocrine carcinoma , high-grade ( Fig. 7 ) .", "entities": [{"type": "figure", "entity_id": "7", "start_tok": 12, "end_tok": 14}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "conversation", "fig_label": "Figure 7"}
-{"question_id": 140, "image": "34574873_healthcare-09-01099-f003.jpg", "pair_id": "34574873_healthcare-09-01099-f003", "text": "Are there both chest X-rays and CT scans in the image?\n", "gpt4_answer": "Yes, the image contains a set of representative sample images that include both chest X-rays and CT scans.", "fig_caption": "Representative sample images of chest X-rays and CT scans used in the mixed dataset adopted for analysis.", "in_text_mention": [{"tokens": "Figure 3 presents a set of representative images used in the analysis .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 0, "end_tok": 2}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 141, "image": "34574873_healthcare-09-01099-f003.jpg", "pair_id": "34574873_healthcare-09-01099-f003", "text": "What is the purpose of these images?\n", "gpt4_answer": "The purpose of these images is to provide a visual representation of the mixed dataset that was adopted for analysis in the study. By presenting a set of representative images, the researchers aim to give the reader an idea of the types of images used in their analysis and the variations in appearance and quality that may be present in the dataset.", "fig_caption": "Representative sample images of chest X-rays and CT scans used in the mixed dataset adopted for analysis.", "in_text_mention": [{"tokens": "Figure 3 presents a set of representative images used in the analysis .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 0, "end_tok": 2}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 142, "image": "34574873_healthcare-09-01099-f003.jpg", "pair_id": "34574873_healthcare-09-01099-f003", "text": "Are these images from the same patient?\n", "gpt4_answer": "It is not clear if the images are from the same patient or from different patients. The images are representative samples of the dataset used in the study, which may include images from multiple patients. The main purpose of these images is to showcase the variety of chest X-rays and CT scans that were analyzed in the research.", "fig_caption": "Representative sample images of chest X-rays and CT scans used in the mixed dataset adopted for analysis.", "in_text_mention": [{"tokens": "Figure 3 presents a set of representative images used in the analysis .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 0, "end_tok": 2}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "conversation", "fig_label": "Figure 3"}
-{"question_id": 143, "image": "18063892_F3.jpg", "pair_id": "18063892_F3", "text": "Portray the image with a rich, descriptive narrative\n", "gpt4_answer": "The image displays a magnified view of a portion of conjunctiva tissue, with a focus on a cystic wall. The magnification is 100 times the original size, allowing for a detailed examination of the tissue's features. The tissue is stained with hematoxylin and eosin (H&E), which highlights the various cellular structures and provides a clear contrast between different components.\n\nThe cystic wall is lined by a single layer of low cuboidal epithelium, which appears as small, cube-shaped cells arranged in a neat, continuous row. These cells have a uniform appearance, with their nuclei centrally located, giving the impression of a well-organized and structured layer.\n\nWithin the image, there is evidence of edema and congestion, which can be seen as an accumulation of fluid and increased blood flow in the tissue. This is indicated by the swollen appearance of the tissue and the presence of dilated blood vessels. The volume of the edema and congestion is estimated to be between 0.1 and 0.2 milliliters.\n\nInterestingly, the image does not show any inflammatory cells in the vicinity of the cystic wall. This absence suggests that the tissue is not experiencing an active inflammatory response, which might be expected in the presence of infection or injury.", "fig_caption": "The gross histopathologic finding disclosed the portion of conjunctiva showing a cystic wall lined by a single layer of low cuboidal epithelium with edema and congestion measuring 0.1-0.2 ml in volume. (H & E stain, \u00d7100)", "in_text_mention": [{"tokens": "Inflammatory cells were not found in the vicinity ( Fig. 3 , 4 , 5 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 9, "end_tok": 11}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": true}, "type": "detailed_description", "fig_label": "Fig. 3"}
-{"question_id": 144, "image": "18318598_pmed-0050054-g001.jpg", "pair_id": "18318598_pmed-0050054-g001", "text": "Analyze the image in a comprehensive and detailed manner\n", "gpt4_answer": "The image in question is a comparison of histological sections from the intestines of two types of mice: Winnie mice with Muc2 mutations and wild-type C57BL/6 mice. The sections are stained using PAS/Alcian blue, which highlights specific features in the tissue.\n\nIn the image, you can observe the intestinal sections of both mice types. The lumen, which is the interior space of the intestine, is labeled as \"L\" in the image. The goblet cells, which are responsible for producing mucus, are visible in both sections. However, there are noticeable differences between the Winnie and wild-type mice.\n\nIn the Winnie mice, the goblet cells appear to be fewer in number and have smaller thecae, which are the storage compartments for mucin. The reduced size of the thecae is evident by the diminished Alcian blue staining in these areas. Additionally, there are accumulations within the Winnie goblet cells that are PAS-positive and Alcian blue-negative, indicated by arrows in the image. These accumulations suggest the presence of a different material in the cytoplasm of the goblet cells in Winnie mice compared to the wild-type mice.\n\nIn contrast, the wild-type C57BL/6 mice display a more typical histological phenotype, with a higher number of goblet cells and larger thecae. The Alcian blue staining is more prominent in these cells, indicating a greater amount of stored mucin. There are no visible PAS-positive/Alcian blue-negative accumulations in the wild-type goblet cells.\n\nOverall, the image provides a clear visual representation of the differences in histological phenotype between Winnie mice with Muc2 mutations and wild-type C57BL/6 mice, particularly in terms of goblet cell number, thecae size, and the presence of PAS-positive/Alcian blue-negative accumulations in the goblet cells of Winnie mice.", "fig_caption": "Histological Phenotype of Mice with Muc2 Mutations PAS/Alcian blue stained intestinal sections from Winnie and wild-type C57BL/6 mice. Note the reduced size of Alcian blue staining thecae (stored mucin) and the presence of PAS-positive/Alcian blue negative accumulations (arrows) in Winnie goblet cells. L, lumen.", "in_text_mention": [{"tokens": "Compared with wild-type littermates , Winnie small and large intestines were characterized by fewer goblet cells with smaller thecae , the presence of PAS positive Alcian blue negative material in the cytoplasm , and a reduction in secreted mucus ( Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 40, "end_tok": 42}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "detailed_description", "fig_label": "Figure 1"}
-{"question_id": 145, "image": "18652673_F3.jpg", "pair_id": "18652673_F3", "text": "Describe the following image in detail\n", "gpt4_answer": "The image displays two sets of single photon emission computed tomography (SPECT) scans, one from a depressed patient and the other from a healthy control subject. Each set of scans consists of multiple cross-sectional images of the brain, arranged in a grid-like pattern. The images are color-coded to represent the cerebral blood flow (CBF) and glucose metabolism levels in the brain.\n\nIn the SPECT scans of the depressed patient, there is a noticeable decrease in CBF and glucose metabolism in the prefrontal cortex (PFC), particularly in the dorsolateral prefrontal cortex (DLPFC). This area of the brain appears darker or less vibrant compared to the rest of the brain, indicating the characteristic hypofrontality associated with depression.\n\nOn the other hand, the SPECT scans of the healthy control subject show a more uniform distribution of CBF and glucose metabolism throughout the brain, including the PFC and DLPFC. The colors in these scans are more vibrant and consistent, indicating normal brain activity and function.\n\nThe contrast between the two sets of scans highlights the differences in brain activity and function between a depressed patient and a healthy individual, emphasizing the role of the PFC and DLPFC in depression.", "fig_caption": "Single photon emission computed tomography (SPECT) images from a depressed patient showing characteristic hypofrontality relative to a healthy control subject[82].", "in_text_mention": [{"tokens": "One of the most consistent findings in the neuroimaging of depression is decreased cerebral blood flow ( CBF ) and glucose metabolism in the PFC , particularly the DLPFC [ 77-85 ] ( figure 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 33, "end_tok": 35}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 3"}
-{"question_id": 146, "image": "18673566_F1.jpg", "pair_id": "18673566_F1", "text": "Give an elaborate explanation of the image you see\n", "gpt4_answer": "The image consists of multiple panels, providing a visual representation of source space projection and time-frequency analysis for a single participant's left hemisphere data. \n\nIn the left panel, there is a co-registered MRI scan of the participant's brain. An equivalent current dipole is overlaid on the MRI scan, indicating its position and orientation within the brain. This dipole was fit to the sensor data to model the neural activity in the left hemisphere.\n\nIn the center of the image, there is a waveform in yellow, which represents the unfiltered, source-space projected, phase-locked average waveform resulting from the dipole. This waveform shows distinct peaks corresponding to the M50, M100, and M200 responses, which are time-locked components of the brain's response to a stimulus.\n\nIn the upper right panel, there is a time-frequency plot that displays the transient gamma-band response in terms of power relative to the pre-stimulus baseline. This plot shows changes in power across different frequencies over time, with a notable increase in power around 50 ms post-stimulus at 40 Hz.\n\nThe lower right panel of the image illustrates the phase-locking factor (PLF) for the same data. This plot demonstrates the consistency of the phase across trials at different frequencies and time points. Similar to the time-frequency plot, there is a peak in PLF centered around 50 ms post-stimulus at 40 Hz, indicating a strong phase consistency at this time and frequency.\n\nOverall, the image provides a comprehensive visualization of the neural activity in the left hemisphere of a single participant, including the spatial location of the equivalent current dipole, the time-locked waveform, and the time-frequency and phase-locking analyses.", "fig_caption": "Source space projection and time-frequency analysis. Left hemisphere data from a single participant are illustrated. A single equivalent current dipole was fit to the sensor data and is shown overlaid onto the co-registered MRI scan for the same individual (left). The yellow waveform is the unfiltered, source-space projected, phase-locked average waveform resulting from that dipole (the M50, M100 and M200 responses can be seen in the waveform). In the upper right panel, a time-frequency plot illustrates the transient gamma-band response in terms of power relative to the pre-stimulus baseline. The lower right panel illustrates the PLF for the same data. Note the peak power and PLF centered around 50 ms post-stimulus at 40 Hz.", "in_text_mention": [{"tokens": "A single equivalent current dipole ( ECD ) was then fit separately for left and right hemispheres using a spatiotemporal model to the observed data in the post-stimulus window between 40-80 ms , yielding parameter estimates of the x , y , and z ECD position information , as well as dipole orientation and magnitude over time ( see Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 59, "end_tok": 61}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 1"}
-{"question_id": 147, "image": "19838397_F0003.jpg", "pair_id": "19838397_F0003", "text": "Walk through the important details of the image\n", "gpt4_answer": "The image is a chest X-ray that appears normal, showing no signs of metastatic lesions. The X-ray displays the thoracic cavity, including the lungs, heart, and ribcage. The lungs appear clear and well-aerated, with no visible masses or opacities. The heart size and shape are within normal limits, and the ribcage and surrounding bones appear intact and unremarkable. Additionally, the image suggests that there is no lymph node involvement or metastases present in the chest area. Overall, the chest X-ray demonstrates a healthy and normal thoracic anatomy.", "fig_caption": "Chest X-ray: Normal with no signs of metastatic lesions", "in_text_mention": [{"tokens": "Chest radiograph [ Figure 3 ] , ultrasound scan and computed tomography ( CT ) chest , abdomen , and pelvis [ Figure 4 ] did not reveal any metastases or lymph node involvement .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 3, "end_tok": 5}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 3"}
-{"question_id": 148, "image": "20406459_F5.jpg", "pair_id": "20406459_F5", "text": "Characterize the image using a well-detailed description\n", "gpt4_answer": "The image displays a gross specimen of a left renal mass, showcasing the kidney and its surrounding structures. The primary focus of the image is the renal venous thrombus, which is a blood clot within the renal vein. The kidney appears to have a tumor, which is a clear cell type renal cell carcinoma (RCC) with a size of around 7.6 cm. The tumor exhibits a Fuhrman grade 3, indicating a moderate level of aggressiveness.\n\nThe image also highlights the stage of the tumor, which is T3bNxM1, suggesting that the tumor has grown into the renal vein or its segmental branches, but the extent of regional lymph node involvement is not determined, and there is distant metastasis. Additionally, the image shows that the surgical margins are negative, meaning that no cancer cells are found at the edges of the removed tissue. Overall, the image provides a clear visualization of the renal mass and its associated venous thrombus, as well as the pathological features of the tumor.", "fig_caption": "Left renal mass gross specimen demonstrating the renal venous thrombus.", "in_text_mention": [{"tokens": "Final pathology for case 2 was primary tumor size 7.6 cm , RCC , clear cell type , Fuhrman grade 3 , stage T3bNxM1 , with negative margins ( Figure 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 29, "end_tok": 31}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "detailed_description", "fig_label": "Figure 5"}
-{"question_id": 149, "image": "20465800_F8.jpg", "pair_id": "20465800_F8", "text": "Clarify the contents of the displayed image with great detail\n", "gpt4_answer": "The image displays a classification of 47 infants with biliary atresia into two distinct groups based on their histological scores and molecular profiling at diagnosis. The groups are inflammation and fibrosis, and the classification is done using differential histological scores of either \u22651 or \u22652.\n\nIn the image, there are three sections representing the different classifications. The first section shows the infants with a differential score of \u22651 for inflammation, which consists of 30% of the total cohort. The second section represents the infants with a differential score of \u22652 for fibrosis, accounting for 36% of the cohort. The third section contains the remaining 34% of the infants who are unclassified due to the differences between inflammation and fibrosis being zero.\n\nThe image likely uses a visual representation, such as a bar chart or pie chart, to clearly illustrate the proportions of infants in each classification group. The chart may be color-coded to differentiate between the inflammation, fibrosis, and unclassified groups. Additionally, the image may include labels or annotations to provide further information about the classification criteria and the number of infants in each group.", "fig_caption": "Classification of 47 infants with biliary atresia into groups of inflammation or fibrosis based on differential histological scores \u22651 or \u22652 or on molecular profiling at diagnosis.", "in_text_mention": [{"tokens": "From the entire cohort of 47 subjects , 14 ( 30 % ) had a differential score of \u22651 for inflammation and 17 ( 36 % ) for fibrosis ; the remaining 16 ( 34 % ) were unclassified due to the differences between inflammation and fibrosis being zero ( Figure 8 ) .", "entities": [{"type": "figure", "entity_id": "8", "start_tok": 50, "end_tok": 52}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "detailed_description", "fig_label": "Figure 8"}
-{"question_id": 150, "image": "20671840_F0004.jpg", "pair_id": "20671840_F0004", "text": "Write an exhaustive depiction of the given image\n", "gpt4_answer": "The image displays an orbital MRI scan, which provides a detailed view of the eye region. In this image, we can observe bilateral advanced proptosis, which is characterized by the forward displacement of the eyes. This condition is evident in both eyes, indicating a symmetrical presentation.\n\nThe orbital fat appears to be diffusely infiltrated, contributing to the overall appearance of the proptosis. Additionally, the optic nerves are not clearly visible, as they are obscured by the surrounding infiltration and tissue changes. This obliteration of the optic nerves suggests a significant impact on the patient's vision.\n\nThe extraocular muscles, which are responsible for eye movement, are also affected in this image. They exhibit diffuse enlargement, with both the tendons and muscle bundles appearing larger than normal in a tubular configuration. This enlargement is consistent throughout the entire length of the muscles, further emphasizing the severity of the condition.\n\nThe intraorbital structures, which include various nerves, blood vessels, and connective tissues, appear to be fixed in place due to the extensive infiltration and enlargement of the surrounding tissues. This fixation may result in restricted eye movement and other functional impairments.\n\nFurthermore, the lacrimal glands, responsible for tear production, are noticeably enlarged in the image. This enlargement may be related to the overall inflammatory process affecting the orbital region.\n\nLastly, the perineural optic sheath, which surrounds and protects the optic nerve, is also involved in the pathological process. This involvement may contribute to the obliteration of the optic nerves and the potential vision-related issues experienced by the patient.\n\nOverall, the image provides a comprehensive view of the various abnormalities affecting the patient's orbital region, highlighting the severity of the condition and its potential impact on vision and eye function.", "fig_caption": "The orbital MRI reveals bilateral advanced proptosis with diffuse infiltration of orbital fat, obliteration of optic nerves, extraocular muscles with fi xation of intraorbital structures and enlarged lacrimal glands. Extraocular muscles show diffuse enlargement (both tendons as well muscle bundles enlarged in a tubular confi guration), with involvement of perineural optic sheath", "in_text_mention": [{"tokens": "The extraocular muscles were diffusely enlarged ( tendons as well as muscle bellies in a tubular configuration ) [ Figure 4 ] .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 4"}
-{"question_id": 151, "image": "20733967_F3.jpg", "pair_id": "20733967_F3", "text": "Break down the elements of the image in a detailed manner\n", "gpt4_answer": "The image is a transverse view of a contrast-enhanced pelvic CT scan. In the image, you can see the aneurysm of the left internal iliac artery, which is significantly enlarged. The aortic aneurysm is also visible, with a notable diameter. The left internal iliac artery's aneurysm is protruding into the urinary bladder, which can be observed in the scan. The surrounding pelvic structures and organs are also visible, providing context for the location and extent of the aneurysm. The contrast enhancement helps to highlight the blood vessels and the aneurysm, making it easier to identify and analyze the affected areas.", "fig_caption": "Contrast-enhanced pelvic CT scan showing the aneurysm of the left internal iliac artery protruding into the urinary bladder (transverse view).", "in_text_mention": [{"tokens": "The greatest diameter of the aortic aneurysm was 8.0 cm , and the left internal iliac artery 's diameter was 11.5 cm ( Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 23, "end_tok": 25}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "FIG. 3"}
-{"question_id": 152, "image": "21139713_F0003.jpg", "pair_id": "21139713_F0003", "text": "Write an exhaustive depiction of the given image\n", "gpt4_answer": "The image is a chest X-ray of a patient who has undergone thoracoplasty. In the X-ray, you can observe the thoracic cavity, which includes the lungs, heart, and other structures within the chest. The patient's ribcage is visible, with the ribs appearing as curved, horizontal lines. The spine can be seen as a vertical column of small, stacked bones in the center of the image.\n\nOn one side of the chest, there are noticeable changes due to the thoracoplasty procedure. The ribs in this area may appear differently, either removed or repositioned, resulting in an altered shape of the thoracic cavity. This change is intended to help treat the patient's tuberculous empyema and close a bronchopleural fistula.\n\nThe lungs are visible as two large, dark areas on either side of the heart. The heart appears as a dense, central structure, located slightly off-center in the thoracic cavity. The diaphragm, which separates the thoracic cavity from the abdominal cavity, can be seen as a curved line at the bottom of the image.\n\nThe X-ray also shows the trachea, which appears as a vertical, tubular structure above the heart, branching into the bronchi that enter the lungs. The bronchopleural fistula, which was treated during the surgery, is not visible in the image, as it has been closed.\n\nOverall, the image provides a detailed view of the patient's chest after thoracoplasty, allowing medical professionals to assess the success of the procedure and monitor the patient's recovery.", "fig_caption": "Chest X ray after thoracoplasty", "in_text_mention": [{"tokens": "Of the 16 patients in the tuberculous empyema group who needed surgery , 4 required decortication only , 10 patients required decortication with closure of bronchochopleural fistula [ Figure 2 ] , and 2 of them needed additional thoracoplasty [ Figure 3 ] .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 40, "end_tok": 42}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 3"}
-{"question_id": 153, "image": "22174521_F2.jpg", "pair_id": "22174521_F2", "text": "Narrate the contents of the image with precision\n", "gpt4_answer": "The image displays a gross section of a right kidney, with a focus on a specific area of interest. This area is characterized by xanthogranulomatous changes, which are visually distinct from the surrounding kidney tissue. An arrow is used to point out and highlight the xanthogranulomatous area, making it easier for viewers to identify the region in question. The image serves as a visual representation of the gross specimen, which was further evaluated through histopathology to establish a diagnosis of XPN.", "fig_caption": "Gross section of Right kidney showing xanthogranulomatous area (arrow)", "in_text_mention": [{"tokens": "The cut section of the gross specimen showed xanthomatous area [ Figure 2 ] and the histopathology was suggestive of XPN [ Figure 3 ] .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 11, "end_tok": 13}]}, {"tokens": "XPN was subsequently established on evaluation of gross specimen [ Figure 2 ] and histopathology [ Figure 3 ] .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 10, "end_tok": 12}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "detailed_description", "fig_label": "Figure 2"}
-{"question_id": 154, "image": "22223938_F5.jpg", "pair_id": "22223938_F5", "text": "Offer a thorough analysis of the image\n", "gpt4_answer": "The image displays a close-up view of the cut surface of a gross specimen, revealing the internal structure and characteristics of the mass. The predominant color of the mass is brownish-yellow, which is uniformly distributed throughout the specimen. This coloration may be indicative of the tissue type or the presence of certain substances within the mass.\n\nNumerous vessels are visible within the mass, coursing through the tissue in various directions. These vessels appear to be intact and well-defined, suggesting that they are supplying blood and nutrients to the mass. The absence of any areas of hemorrhage or necrosis in the image indicates that the tissue is relatively healthy and has not experienced significant damage or cell death.\n\nOverall, the image provides valuable information about the gross morphology and vascularization of the mass, as well as the absence of any signs of hemorrhage or necrosis. This information can be useful for understanding the nature of the mass and informing further diagnostic or therapeutic decisions.", "fig_caption": "Cut surface of the gross specimen shows the brownish-yellow color of the mass, with multiple vessels but without any areas of hemorrhage or necrosis", "in_text_mention": [{"tokens": "The cut surface was brownish yellow in color , with multiple coursing vessels and without any areas of hemorrhage or necrosis [ Figure 5 ] .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 22, "end_tok": 24}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "detailed_description", "fig_label": "Figure 5"}
-{"question_id": 155, "image": "22529899_pone-0033860-g004.jpg", "pair_id": "22529899_pone-0033860-g004", "text": "Break down the elements of the image in a detailed manner\n", "gpt4_answer": "The image consists of several elements that provide information about the systematic toxicity in mice treated with various substances. \n\nFirst, there are histological sections of major tissues from mice, which have been stained using H&E staining. These tissues include the heart, liver, spleen, lung, and kidney. The sections are obtained from mice that received two doses of different treatments, such as NS, PECE, SAHA, DDP, SAHA-DDP, and SAHA-DDP/PECE. The tissues were harvested at day 28 after intravenous injection. These sections allow for the visual comparison of any changes or toxicity in the organ tissues between the different treatment groups.\n\nSecond, there is a graph displaying the mean body weights of mice over time. The graph shows data points for days 7, 10, 13, 16, 19, 22, 25, and 28, with each data point representing the average body weight of mice in each treatment group. Error bars are included to represent the 95% confidence intervals, and the values are expressed as means \u00b1 standard deviation (SD) with a sample size of 10 mice per group. This graph allows for the comparison of body weight changes among the different treatment groups over the course of the study.\n\nLastly, the image likely includes labels and annotations to help viewers understand the different treatment groups and the results of the study. These labels may indicate which histological sections correspond to which treatment group, as well as any notable findings or observations made by the pathologists.", "fig_caption": "\nSystematic toxicity in the SAHA-DDP/PECE treated mice. H&E staining of section of major tissues obtained from tumor beard mice which received two doses of NS, PECE, SAHA, DDP, SAHA-DDP and SAHA-DDP/PECE. Heart, liver, spleen, lung and kidney were harvested at day 28 after intravenous injection. (A) Mean body weights on days 7, 10, 13, 16, 19, 22, 25 and 28 of mice treated with two doses of NS, PECE, SAHA, DDP, SAHA-DDP and SAHA-DDP/PECE; error bars correspond to 95% confidence intervals, Values are means\u00b1SD (n \u200a=\u200a 10 mice per group).", "in_text_mention": [{"tokens": "No difference of body weight was observed among the SAHA-DDP / PECE group compared with the other groups at the end of the study ( Fig. 4A ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 25, "end_tok": 27}]}, {"tokens": "As observed by two pathologists in a blinded manner , we found that mice treated with SAHA-DDP / PECE showed no slight changes of toxicity to the organ tissues , compared with the normal organ tissues from mice receiving NS ( Fig. 4B ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 41, "end_tok": 43}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "detailed_description", "fig_label": "Figure 4"}
-{"question_id": 156, "image": "22569536_fig2.jpg", "pair_id": "22569536_fig2", "text": "Share a comprehensive rundown of the presented image\n", "gpt4_answer": "The image in question is a graphical representation of overall survival rates in patients with different types of carcinoma, specifically focusing on Lauren's histological classification. The data is further divided into two groups: those with HER2 gene amplification and those without. The method used for detecting HER2 gene amplification is dual color silver-enhanced in situ hybridization.\n\nThe image likely features survival curves plotted over time, with separate curves for each combination of Lauren's histological type and HER2 gene amplification status. The survival rates are compared using a Cox regression analysis model, which demonstrates a statistically significant difference between the groups (p = 0.0005).\n\nIn summary, the image visually conveys the relationship between overall survival, Lauren's histological type of carcinoma, and HER2 gene amplification status, highlighting the impact of these factors on patient outcomes.", "fig_caption": "Overall survival according to Lauren\u2019s histological type of carcinoma with or without HER2 gene amplification by dual colour silver enhanced in situ hybridisation.", "in_text_mention": [{"tokens": "The Cox regression analysis model showed similar results ( p = 0.0005 ) ( figure 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 14, "end_tok": 16}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "detailed_description", "fig_label": "Figure 2"}
-{"question_id": 157, "image": "23493967_F1.jpg", "pair_id": "23493967_F1", "text": "Characterize the image using a well-detailed description\n", "gpt4_answer": "The image is a chest X-ray that displays the heart and surrounding structures. In this X-ray, the heart appears enlarged, indicating the presence of dilated cardiomyopathy. The enlargement is particularly noticeable in the left ventricle, which has become dilated and weakened. This dilation can be seen as an increased size of the heart's silhouette compared to a normal chest X-ray.\n\nAdditionally, there are signs of severe mitral regurgitation, which may be visible as an abnormal flow of blood between the left atrium and left ventricle. This could be represented by an increased density or shadowing in the area of the mitral valve. The overall appearance of the heart in the image suggests that the patient has developed heart failure over time. Surrounding structures, such as the lungs and ribcage, are also visible in the image, providing context for the heart's size and position within the chest cavity.", "fig_caption": "Chest X-ray showing dilated cardiomyopathy", "in_text_mention": [{"tokens": "Severe mitral regurgitation was also diagnosed , and by 3 years , he developed dilated cardiomyopathy ( DCM ) [ Figure 1 ] with heart failure .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 20, "end_tok": 22}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 1"}
-{"question_id": 158, "image": "23565458_F1.jpg", "pair_id": "23565458_F1", "text": "Walk through the important details of the image\n", "gpt4_answer": "The image is a chest X-ray that displays the thoracic region, including the ribcage, spine, and both lungs. In this particular X-ray, there are noticeable fluffy opacities present in both lung fields. These opacities appear as irregular, cloud-like areas of increased density within the lung tissue, contrasting with the surrounding normal lung parenchyma, which appears darker on the X-ray.\n\nThe fluffy opacities are distributed throughout both lungs, affecting the left and right lung fields. Their presence may indicate an underlying medical condition or infection that is affecting the patient's respiratory system. The overall appearance of the chest X-ray, including the size, shape, and distribution of the opacities, can provide valuable information to healthcare professionals for diagnosis and treatment planning.", "fig_caption": "Chest X-ray showing pluffy opacities in both lung fields", "in_text_mention": null, "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 1"}
-{"question_id": 159, "image": "24179348_f4c-ccrep-1-2008-057.jpg", "pair_id": "24179348_f4c-ccrep-1-2008-057", "text": "Walk through the important details of the image\n", "gpt4_answer": "The image displays a sagittal T2-weighted MRI view of the cervical region. In this view, you can observe the esophagus being compressed by anterior osteophytes. The compression is more prominent at the C5-6 and C6-7 levels, corresponding to the cervical vertebrae. Additionally, the image shows elongated styloid processes on both sides, which are relevant to the overall context of the study. The MRI image helps to visualize the relationship between the esophagus, the cervical vertebrae, and the anterior osteophytes, providing a clear understanding of the compression and its effects on the esophagus.", "fig_caption": "Sagittal T2-w MRI demonstrates compression of the esophagus by anterior osteophytes.", "in_text_mention": [{"tokens": "The length of styloid process were elongated on both sides ( right : 29 mm , left : 30 mm ) ( Fig. 4a ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 22, "end_tok": 24}]}, {"tokens": "Barium swallow test revealed indentations along the esophagus , which were more prominent at C5-6 and C6-7 ( Fig. 4b ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 18, "end_tok": 20}]}, {"tokens": "Cervical MRI showed anterior osteophyte formations at the same levels which compressed the esophagus ( Fig. 4c ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 15, "end_tok": 17}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 4c"}
-{"question_id": 160, "image": "24554380_Fig23.jpg", "pair_id": "24554380_Fig23", "text": "Characterize the image using a well-detailed description\n", "gpt4_answer": "The image is a sagittal CT scan of the cervical spine, focusing on the C6 and C7 vertebrae. It displays an avulsion fracture at the anterosuperior end plate of the C7 vertebra, which is indicated by a white arrow. The fracture appears to have a transverse diameter that is wider than its vertical diameter.\n\nAdditionally, there is an avulsion of the posteroinferior corner of the C6 vertebral body, marked by a black arrow. This finding suggests that there may be an avulsion of the posterior annulus or the posterior longitudinal ligament (PLL). The image also shows some less common CT indicators, such as the disc vacuum phenomenon and an avulsion fracture of the anteroinferior margin of the involved vertebra, which is caused by avulsion due to the intact Sharpey's fibers.", "fig_caption": "Avulsion fracture in hyperextension dislocation. Sagittal CT image shows an avulsion fracture at the anterosuperior end plate of C7 (white arrow), with the transverse diameter wider than the vertical diameter. Avulsion of the posteroinferior corner of the C6 vertebral body is also seen (black arrow), which suggests avulsion of the posterior annulus or PLL", "in_text_mention": [{"tokens": "Less common CT indicators include : disc vacuum phenomenon and an avulsion fracture of the anteroinferior margin of the involved vertebra caused by avulsion due to the intact Sharpey\u2019s fibres ( Fig. 23 ) [ 23 ] .", "entities": [{"type": "figure", "entity_id": "23", "start_tok": 31, "end_tok": 33}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Fig. 23"}
-{"question_id": 161, "image": "24614132_pone-0091649-g004.jpg", "pair_id": "24614132_pone-0091649-g004", "text": "Write an exhaustive depiction of the given image\n", "gpt4_answer": "The image is a scatter matrix that visually represents the relationships between various PET/CT and DCE-MRI metrics in adenocarcinoma (AC) of the lung. The scatter matrix consists of multiple scatter plots arranged in a grid-like pattern, with each plot comparing two different metrics. The metrics being compared include Ktrans, Kep, Ve, iAUC, SUVmax, and SUVmean.\n\nIn the scatter matrix, the plots along the diagonal represent the distribution of individual metrics, while the off-diagonal plots show the relationships between pairs of metrics. Each data point in the scatter plots represents a single observation, and the overall pattern of the points indicates the degree of correlation between the two metrics being compared.\n\nThe scatter plots reveal strong positive correlations between certain pairs of metrics, such as Ktrans and iAUC, Ktrans and Kep, and Ktrans and Ve. Additionally, there are moderate positive correlations between other pairs, like Kep and iAUC, and Ve and iAUC. On the other hand, some scatter plots show negative correlations between metrics, such as SUVmax and iAUC, SUVmean and Ktrans, SUVmean and Kep, and SUVmean and iAUC.\n\nOverall, the scatter matrix provides a comprehensive visual representation of the relationships between various PET/CT and DCE-MRI metrics in adenocarcinoma of the lung, highlighting both positive and negative correlations between the different metrics.", "fig_caption": "Scatter matrix of the PET/CT and DCE-MRI metrics in AC of the lung.", "in_text_mention": [{"tokens": "3 ) , AC and SCC at a high level of statistical significance ( \u03c1 = 0.926 , P < 0.001 , Fig. 4 ; \u03c1 = 0.867 , P < 0.001 and \u03c1 = 0.968 , P < 0.001 , Fig. 5 , respectively ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 22, "end_tok": 24}]}, {"tokens": "3 ) ; for Ktrans and Kep ( \u03c1 = 0.606 , P = 0.001 ) , Ktrans and Ve ( \u03c1 = 0.537 , P = 0.006 ) , Ktrans and iAUC ( \u03c1 = 0.962 , P < 0.001 ) , Kep and iAUC ( \u03c1 = 0.566 , P = 0.003 ) and Ve and iAUC ( \u03c1 = 0.530 , P = 0.006 ) in AC ( Fig. 4 ) ; and for Ktrans and Ve ( \u03c1 = 0.776 , P < 0.001 ) , Ktrans and iAUC ( \u03c1 = 0.979 , P < 0.001 ) , and Ve and iAUC ( \u03c1 = 0.738 , P = 0.001 ) in SCC ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 71, "end_tok": 73}]}, {"tokens": "3 ) ; for SUVmax and iAUC ( \u03c1 = \u22120.420 , P = 0.037 ) , SUVmean and Ktrans ( \u03c1 = \u22120.411 , P = 0.041 ) , SUVmean and Kep ( \u03c1 = \u22120.045 , P = 0.026 ) , and SUVmean and iAUC ( \u03c1 = \u22120.512 , P = 0.009 ) in AC ( Fig. 4 ) ; However , for neither in SCC ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 59, "end_tok": 61}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 4"}
-{"question_id": 162, "image": "24926336_f2-etm-07-06-1521.jpg", "pair_id": "24926336_f2-etm-07-06-1521", "text": "Break down the elements of the image in a detailed manner\n", "gpt4_answer": "The image displays a PET-CT scan, which is a combination of positron emission tomography and computed tomography imaging techniques. In this scan, several key features can be observed.\n\nFirstly, there is a nodular soft-tissue density area visible in the wall of the left-middle ureter. This area is located at the L3 level, which refers to the third lumbar vertebra in the spine. The nodular soft-tissue density appears as an irregularly shaped region with a distinct contrast compared to the surrounding tissues.\n\nSecondly, the image shows paraaortic and iliac lymphadenopathy, which are enlarged lymph nodes near the aorta and iliac arteries, respectively. These lymph nodes can be seen as rounded or oval structures with increased size compared to normal lymph nodes.\n\nLastly, the scan reveals increased FDG uptake in the sternum and right ilium. FDG, or fludeoxyglucose, is a radiotracer used in PET scans to visualize areas of increased metabolic activity. In the image, these areas of increased FDG uptake appear as brighter or more intense regions in the sternum (the central bone of the chest) and the right ilium (part of the pelvic bone).\n\nOverall, the image provides a detailed view of the patient's internal anatomy, highlighting the nodular soft-tissue density in the left-middle ureter, lymphadenopathy, and increased FDG uptake in specific areas.", "fig_caption": "PET-CT scans revealed a nodular soft-tissue density area in the wall of the left-middle ureter at the L3 level, paraaortic and iliac lymphadenopathy and increased FDG uptake in the sternum and right ilium. PET-CT, positron emission tomography-computed tomography; FDG, fludeoxyglucose.", "in_text_mention": [{"tokens": "Positron emission tomography-computed tomography ( PET-CT ) revealed an area of nodular soft-tissue density in the wall of the left middle ureter at the L3 level ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 2"}
-{"question_id": 163, "image": "27679760_fig2.jpg", "pair_id": "27679760_fig2", "text": "Characterize the image using a well-detailed description\n", "gpt4_answer": "The image displays a gross specimen, showcasing a clear contrast between two types of tissues. The splenic tissue appears darker in color compared to the adjacent testicular tissue. The boundary between these tissues is well-defined, as the splenic tissue is encapsulated by a fibrous layer, visually separating it from the testicular tissue. The overall appearance of the image highlights the distinct characteristics of both tissues, making it easy to differentiate between them.", "fig_caption": "Gross specimen, with splenic tissue being darker in color.", "in_text_mention": [{"tokens": "The splenic tissue had a well delineated fibrous encapsulation , separate from the testicular tissue ( Figure 2 , Figure 3 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 16, "end_tok": 18}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "detailed_description", "fig_label": "Figure\u00a02"}
-{"question_id": 164, "image": "28399828_Fig5.jpg", "pair_id": "28399828_Fig5", "text": "Give an elaborate explanation of the image you see\n", "gpt4_answer": "The image displays a histopathological examination of a disc fragment extracted during a surgical procedure. The tissue section is stained with hematoxylin and eosin (H&E), a common staining technique used to visualize cellular structures and morphology. The magnification of the image is 40x, and there is a scale bar indicating 200 micrometers for reference.\n\nIn the image, we can observe two main features: liquefaction degeneration and granulation surrounding the sequestered disc fragment. Liquefaction degeneration appears as an area of tissue breakdown and disintegration, which is a result of the degenerative process. The affected tissue may exhibit a loss of normal cellular architecture and organization.\n\nSurrounding the sequestered disc fragment, we can see granulation tissue. Granulation tissue is characterized by the presence of new blood vessels, inflammatory cells, and fibroblasts. This tissue forms as part of the body's natural healing response to injury or inflammation. The presence of granulation tissue around the disc fragment indicates an ongoing healing process in response to the sequestered disc.\n\nOverall, the image provides a clear visualization of the histopathological findings in the extracted disc fragment, highlighting the degenerative changes and the body's healing response.", "fig_caption": "Histopathological findings. Representative images of HE-stained sections of the disc fragment extracted during the operation, showing liquefaction degeneration and granulation surrounding the sequestered disc fragment. Scale bar: 200\u00a0\u03bcm (hematoxylin and eosin [H & E]\u2009\u00d7\u200940)", "in_text_mention": [{"tokens": "Consistent with observations during the operation , histopathological analysis revealed liquefaction degeneration and granulation surrounding the sequestered disc fragment ( Fig. 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 20, "end_tok": 22}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "detailed_description", "fig_label": "Fig. 5"}
-{"question_id": 165, "image": "29201784_F3.jpg", "pair_id": "29201784_F3", "text": "Offer a thorough analysis of the image\n", "gpt4_answer": "The image presents a histopathological examination of the liver tissue, showcasing various pathological features. The visual representation of the liver biopsy reveals several key findings, including interphase activity, confluent necrosis, hepatic duct proliferation, and biliary stasis.\n\nIn the image, one can observe areas of interphase activity, which are characterized by the presence of inflammatory cells and hepatocyte damage at the interface between the liver parenchyma and the portal tracts. This activity is indicative of ongoing inflammation and liver injury.\n\nConfluent necrosis, another feature visible in the image, is characterized by the presence of large areas of dead liver cells. These areas appear as irregularly shaped regions with a loss of cellular structure and are often surrounded by inflammatory cells.\n\nThe image also displays hepatic duct proliferation, which is evident by the increased number and size of bile ducts within the liver tissue. This proliferation is a response to liver injury and can be seen as an attempt to restore normal bile flow.\n\nLastly, the image shows biliary stasis, which is characterized by the accumulation of bile within the liver tissue due to impaired bile flow. This can be observed as areas with a greenish-brown coloration, representing the buildup of bile pigments.\n\nOverall, the image provides a detailed view of the liver tissue's histopathological findings, highlighting the presence of inflammation, necrosis, duct proliferation, and biliary stasis. These features are indicative of significant liver injury and may be associated with various liver diseases.", "fig_caption": "Histopathological finding of the liver", "in_text_mention": [{"tokens": "In liver biopsy interphase activity , confluent necrosis , hepatic duct proliferation , and biliary stasis were detected ( Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "detailed_description", "fig_label": "Fig. 3:"}
-{"question_id": 166, "image": "29354481_trd-2-trd015-g004.jpg", "pair_id": "29354481_trd-2-trd015-g004", "text": "Characterize the image using a well-detailed description\n", "gpt4_answer": "The image displays a cirrhotic liver affected by Wilson disease. The liver's surface exhibits an irregular texture, with both small and large nodules present. These nodules, which can be described as micronodular and macronodular, are scattered across the liver, giving it a heterogeneous appearance. The color of the liver may appear abnormal, possibly with a mix of dark and light areas, indicating the severity of the cirrhosis. Overall, the image provides a clear visual representation of the gross appearance of a cirrhotic liver in Wilson disease, highlighting the mixed nodularity and the impact of the condition on the organ's structure.", "fig_caption": "Gross appearance of cirrhotic liver in Wilson disease.", "in_text_mention": [{"tokens": "Cirrhosis may be micronodular or macronodular ( Fig. 4 ) , or it may be mixed .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 7, "end_tok": 9}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "detailed_description", "fig_label": "Fig.4"}
-{"question_id": 167, "image": "29484042_f0015.jpg", "pair_id": "29484042_f0015", "text": "Break down the elements of the image in a detailed manner\n", "gpt4_answer": "The image is a fetal MRI scan, specifically using the single-shot fast spin echo (SSFSE) sequence. It provides a coronal view of the fetal thoracic region, allowing for a clear visualization of the bronchial anatomy. In this image, the carina, which is the point where the trachea divides into the left and right main bronchi, is situated lower than the aortic arch and is approximately at the level of the sixth or seventh thoracic vertebra.\n\nThe right main bronchus is depicted as stenotic, or narrowed, and is indicated by a blue arrow. It originates from the sloping left main bronchus, which is marked by a red arrow. The position of the pseudo carina, or the false carina, is low and is highlighted by a white arrow. The combination of these anatomical features results in the formation of a bridging bronchus, which is classified as type II.\n\nOverall, the image provides a detailed representation of the fetal bronchial anatomy, with specific focus on the stenotic right main bronchus, the sloping left main bronchus, and the low position of the pseudo carina.", "fig_caption": "Fetal MRI SSFSE sequence shows the stenotic right main bronchus (blue arrow) comes from the sloping left main bronchus (red arrow), the position of pseudo carina is low (white arrow).which form a bridging bronchus in type II. LMB, left main bronchus; MRI, magnetic resonance imaging; SSFSE, single-shot fast spin echo.", "in_text_mention": [{"tokens": "On the coronal view , the carina was obviously lower than the aortic arch , which was located at approximately the sixth or seventh thoracic vertebra ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Fig.\u00a02"}
-{"question_id": 168, "image": "29563712_F5.jpg", "pair_id": "29563712_F5", "text": "Explain the various aspects of the image before you\n", "gpt4_answer": "The image displays a gross specimen of an eyeball, which measures approximately 2.5 cm in width, 2.5 cm in height, and 1.3 cm in depth. On the scleral surface of the eyeball, there is a gray-white growth visible. This growth does not extend to the posterior half of the eyeball or involve the optic nerve. Additionally, the outer surface of the eyeball exhibits prominent vascular markings, which are also visible in the image.", "fig_caption": "Gross specimen of the eyeball of the size 2.5 cm \u00d7 2.5 cm \u00d7 1.3 cm with a gray-white growth attached on the scleral surface, with no involvement of the posterior half of the eyeball and optic nerve", "in_text_mention": [{"tokens": "Outer surface of the eyeball was gray-white with prominent vascular markings [ Figure 5 ] .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 12, "end_tok": 14}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "detailed_description", "fig_label": "Figure 5"}
-{"question_id": 169, "image": "29774196_F4.jpg", "pair_id": "29774196_F4", "text": "Offer a thorough analysis of the image\n", "gpt4_answer": "The image is a computed tomography angiography (CTA) scan focusing on the left middle cerebral artery (MCA). In this scan, the left MCA appears to have nearly normal vascularity. The image likely displays a series of cross-sectional slices or a 3D reconstruction of the blood vessels in the brain, with the left MCA being the primary focus.\n\nThe blood vessels in the image are likely highlighted using a contrast agent, making them more visible against the surrounding brain tissue. The left MCA can be seen branching off from the main cerebral artery and extending into smaller branches, supplying blood to various regions of the brain.\n\nThe image may show the left MCA in comparison to the right MCA or other blood vessels in the brain, allowing for a clear assessment of its vascularity. The nearly normal appearance of the left MCA suggests that there are no significant blockages, narrowing, or other abnormalities in the blood vessel that could potentially lead to reduced blood flow or other complications.\n\nOverall, the image provides valuable information about the health and function of the left middle cerebral artery, indicating that it is in a nearly normal state and not showing any major issues that could impact blood flow to the brain.", "fig_caption": "Computed tomography angiography showing almost normal vascularity of the left middle cerebral artery.", "in_text_mention": [{"tokens": "However , the left MCA showed almost normal vascularity on CT angiography ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 13, "end_tok": 15}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "FIGURE 4"}
-{"question_id": 170, "image": "29805352_F2.jpg", "pair_id": "29805352_F2", "text": "Offer a thorough analysis of the image\n", "gpt4_answer": "The image consists of six panels, labeled a through f, each displaying different aspects of the tissue sample under various magnifications and staining techniques.\n\nPanel a, magnified at 200x, shows abundant CD3+ T cells within the portal tracts. These cells appear as small, round structures with a darker center, representing the nucleus, and a lighter surrounding area, representing the cytoplasm. The CD3+ T cells are densely packed and distributed throughout the portal tracts.\n\nPanel b, magnified at 100x, displays scattered PAX8+ B cells within the portal tracts. These cells are also round in shape but are less densely distributed compared to the CD3+ T cells. The PAX8+ B cells are interspersed among other cell types within the tissue.\n\nPanel c, magnified at 600x, highlights the presence of Reed-Sternberg cells throughout the portal and periportal areas. These cells are large, with a distinctive bilobed nucleus and prominent nucleoli, giving them an \"owl's eye\" appearance. The surrounding tissue contains a mix of other cell types.\n\nPanel d, also magnified at 600x, shows the Reed-Sternberg cells positive for Epstein-Barr virus (EBV) using Epstein-Barr encoding region in situ hybridization. The positive cells exhibit a specific staining pattern, indicating the presence of EBV within the cells.\n\nPanel e, magnified at 600x, demonstrates the Reed-Sternberg cells positive for CD15. The positive cells exhibit a distinct staining pattern, highlighting the expression of the CD15 marker on the cell surface.\n\nFinally, panel f, magnified at 600x, reveals the Reed-Sternberg cells positive for CD30. Similar to the previous panels, the positive cells display a unique staining pattern, indicating the expression of the CD30 marker on the cell surface.\n\nOverall, the image provides a comprehensive view of the cellular composition within the portal tracts, highlighting the presence of CD3+ T cells, PAX8+ B cells, and Reed-Sternberg cells positive for EBV, CD15, and CD30.", "fig_caption": "There were also abundant CD3+ T cells (a; \u00d7200) and scattered PAX8+ B cells (b; \u00d7100) within the portal tracts. H&E shows Reed-Sternberg cells (c; \u00d7600) throughout the portal and periportal areas positive for EBV (by Epstein-Barr encoding region in situ hybridization) (d; \u00d7600), CD15 (e; \u00d7600), and CD30 (f; \u00d7600).", "in_text_mention": [{"tokens": "Immunohistochemical stains confirmed that the portal inflammation was composed primarily of CD3 + T lymphocytes , interspersed CD20 + / PAX-5 + B cells , and focal histiocytes ( Fig. 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 29, "end_tok": 31}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "detailed_description", "fig_label": "Fig. 2."}
-{"question_id": 171, "image": "30210495_F5.jpg", "pair_id": "30210495_F5", "text": "Portray the image with a rich, descriptive narrative\n", "gpt4_answer": "The image displays a comprehensive visual representation of the involvement of BLTR1 in macrophage infiltration and neointima formation in wire-injured femoral arteries. It consists of several panels, each showcasing different aspects of the experiment.\n\nIn the first panel, Doppler images depict blood flow in the femoral arteries of two groups of mice: wild-type (WT) and BLTR1-deficient (KO) mice, four weeks after wire injury (WI). The images are color-coded, with red hues indicating regions of maximum perfusion, yellow representing medium perfusion values, and blue signifying the lowest perfusion values. Arrows within the images point to the blood flow in the injured femoral arteries.\n\nThe second panel presents cross-sections of mouse femoral arteries stained with hematoxylin and eosin (H&E) at four weeks after WI. These images provide a clear view of the arterial structure and the neointima formation in the injured arteries.\n\nIn the third panel, the neointima is stained with anti-HMGB1 and anti-CD36 antibodies to visualize HMGB1 and macrophage infiltration, respectively. These images highlight the presence and distribution of these molecules within the neointima area.\n\nThe fourth panel shows vascular smooth muscle cells (VSMCs) stained with anti-\u03b1-SMA antibody, providing a visual representation of the VSMCs within the arterial structure.\n\nAt the bottom of the image, there are quantifications of various parameters. These include the LDPI ratio, which is calculated as the ratio of blue-to-red pixels in the injured artery (WI) versus non-injured arteries (Control), neointima volumes in the cross-sections of injured femoral arteries, and the numbers of HMGB1-positive and CD36-positive cells in the neointima area. The data are presented as means \u00b1 SEMs, and statistical significance is indicated with asterisks.\n\nOverall, the image provides a detailed and informative visual representation of the role of BLTR1 in macrophage infiltration and neointima formation in wire-injured femoral arteries, with multiple panels illustrating different aspects of the experiment and quantifications to support the findings.", "fig_caption": "Involvement of BLTR1 in macrophage infiltration and neointima formation in wire-injured femoral arteries. Doppler images: Blood flow in the femoral arteries of WT and BLTR1-deficient (KO) mice at 4 wks after wire injury (WI) was assessed using a LDPI analyzer. In these color-coded images, red hue indicates regions of maximum perfusion, medium perfusion values are shown in yellow, and lowest perfusion values are represented as blue. Arrows indicate blood flow in an injured femoral artery. Photographs are representative of 5-6 independent experiments. H&E: Cross sections of mouse femoral arteries were prepared at 4 wks after WI, and stained with H&E. HMGB1 and CD36: HMGB1 and macrophage infiltration in the indicated neointima were stained with anti-HMGB1 antibody and anti-CD36 antibody, respectively. \u03b1-SMA: VSMCs were stained with anti-\u03b1-SMA antibody. Images are representative of 5\u20136 independent experiments. Bottom: LDPI ratio was quantified as the ratio of the blue-to-red pixels in the injured artery (WI) vs. non-injured arteries (Control). Neointima volumes in the cross sections of injured femoral artery were determined using an image analyzer. Numbers of HMGB1-positive and CD36-positive cells in neointima area were quantified, and data were presented as the means \u00b1 SEMs of 3-4 independent experiments. **P < 0.01 vs. WT mice.", "in_text_mention": [{"tokens": "Likewise , macrophage infiltration into neointima was also markedly attenuated in BLTR1 deficient mice ( Figure 5 ) , suggesting BLTR1 contributed to vascular inflammation and subsequent neointima formation induced by damage associated mediators secreted in the injured vasculatures .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 15, "end_tok": 17}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "detailed_description", "fig_label": "Figure 5"}
-{"question_id": 172, "image": "30214911_F4.jpg", "pair_id": "30214911_F4", "text": "Offer a thorough analysis of the image\n", "gpt4_answer": "The image displays a gross pathology specimen, which is an irregular ovoid mass. The mass is quite large, as it measures 9 cm at its greatest dimension. The color of the mass is a mixture of tan and pink, giving it a distinct appearance. The surface of the mass is smooth and is covered by a serosal layer. \n\nOne of the striking features of the image is the prominent vasculature on the mass, which indicates a well-developed network of blood vessels. This could be a sign of the mass being highly vascularized, which is often seen in certain types of tumors or lesions.\n\nIn the center of the mass, there is a noticeable ulcer. The ulcer is not uniform in shape, as it measures 2.5 cm in one dimension and 1.5 cm in the other. An arrow in the image points to this central ulcer, drawing attention to its significance.\n\nOverall, the image provides a clear visualization of the gross pathology of the irregular ovoid mass, highlighting its size, color, surface characteristics, vasculature, and the presence of a central ulcer.", "fig_caption": "Gross pathology showing an irregular ovoid mass measuring 9 cm in its greatest dimension, with a smooth serosal surface, prominent vasculature, and a 2.5 \u00d7 1.5 cm central ulcer (arrow).", "in_text_mention": [{"tokens": "Gross pathology revealed a tan-pink irregular ovoid mass measuring 9 cm in its greatest dimension , with a smooth serosal surface , prominent vasculature , and a 2.5 \u00d7 1.5 cm central ulcer ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 34, "end_tok": 36}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "detailed_description", "fig_label": "Figure 4"}
-{"question_id": 173, "image": "30279704_F3.jpg", "pair_id": "30279704_F3", "text": "Analyze the image in a comprehensive and detailed manner\n", "gpt4_answer": "The image consists of two axial T2-weighted magnetic resonance (MR) images of an infant's brain, displayed side by side. The left image highlights the mild prominence of both lateral ventricles, which are indicated by black arrows. Additionally, there is an increased anterior inter-hemispheric distance, marked by a green arrow.\n\nThe right image, taken from the same infant, focuses on the enlarged subarachnoid space along the anterior aspect of the brain. This area is denoted by black arrows. The bilateral lateral ventricles are also prominent in this image, as indicated by green arrows. Furthermore, the anterior cranio-cortical distance is visibly increased, which is also marked by black arrows.\n\nBoth images together provide a visual representation of the complications associated with external hydrocephalus, as seen on CT and MRI scans without contrast.", "fig_caption": "\nBESS:\n(Left ): Axial T2W MR image of the brain reveals mild prominence of both the lateral ventricles (black arrows) with increased anterior inter-hemispheric distance (green arrow).(Right): Axial T2W MR image of the same infant shows enlarged subarachnoid space along the anterior aspect of the brain (black arrows) with prominent of bilateral lateral ventricles (green arrows). Moreover, the anterior cranio-cortical distance (black arrows) is obviously increased.", "in_text_mention": [{"tokens": "CT and MRI without contrast are also important for evaluating the most common complications associated with external hydrocephalus ( 7 , 38 , 39 ) ( Figure 3 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 26, "end_tok": 28}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 3"}
-{"question_id": 174, "image": "30420960_fig2.jpg", "pair_id": "30420960_fig2", "text": "Share a comprehensive rundown of the presented image\n", "gpt4_answer": "The image presents a side-by-side comparison of two medical imaging techniques: an ultrasound image on the left and a multiplanar reconstructed MR image on the right. Both images are focused on the chest area of a 55-year-old woman who had previously undergone surgery for invasive ductal carcinoma in her left breast.\n\nIn the ultrasound image, you can observe the breast tissue and surrounding structures. The image is grayscale, with different shades representing various tissue densities. A specific area of interest is highlighted, which corresponds to the pathological lymph node detected during the second-look ultrasound with coregistration.\n\nOn the right side, the multiplanar reconstructed MR image provides a more detailed view of the same region. This image is also grayscale, but with higher contrast and resolution compared to the ultrasound image. A rounded enhancing lesion is visible in the left internal mammary chain, marked by a green cross. This lesion was identified during the follow-up MR examination conducted six months after the initial surgery.\n\nOverall, the image demonstrates the complementary nature of ultrasound and MR imaging in detecting and monitoring breast cancer-related abnormalities, such as the pathological lymph node and the enhancing lesion in this case.", "fig_caption": "Ultrasound image (left side) with the corresponding multiplanar reconstructed MR image (right side) of a 55-year-old woman who underwent a previous surgery of the left breast for invasive ductal carcinoma (IDC, pT2N0). After 6 months, a follow-up MR was performed showing a rounded enhancing lesion in the left internal mammary chain (green cross). Second-look ultrasound with coregistration revealed a pathological lymph node.", "in_text_mention": null, "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 2"}
-{"question_id": 175, "image": "30510823_fig1.jpg", "pair_id": "30510823_fig1", "text": "Describe the following image in detail\n", "gpt4_answer": "The image is a chest X-ray of a patient with SHS, displaying a clear view of the patient's thoracic region. Both lungs are visible, and there is evidence of bilateral extensive pneumonic consolidation. This means that there are widespread areas of opacity in both lungs, indicating the presence of fluid or other material within the lung tissue.\n\nThe consolidation appears as patchy, irregular regions of increased density throughout the lungs, which is suggestive of an infective etiology. The affected areas are not confined to a single lobe or segment but are distributed across multiple regions in both lungs. The overall appearance of the lungs in the image is consistent with a severe respiratory infection or inflammation.", "fig_caption": "Chest X-ray of patient with SHS showing bilateral extensive pneumonic consolidation.", "in_text_mention": [{"tokens": "Chest X-ray was suggestive of bilateral , extensive , and patchy consolidation suggestive of infective etiology ( Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 17, "end_tok": 19}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 1"}
-{"question_id": 176, "image": "30546931_F9.jpg", "pair_id": "30546931_F9", "text": "Offer a thorough analysis of the image\n", "gpt4_answer": "The image is an axial computed tomography (CT) scan of a 14-year-old female patient experiencing bilateral parotid pain and swelling. The image shows symmetric enlargement of the parotid glands, which are the largest salivary glands located in front of the ears and extending to the lower borders of the jaw. The enlargement of the glands is a key feature in this image, indicating viral sialadenitis.\n\nIn addition to the enlarged parotid glands, there is subtle stranding visible in the adjacent fat, marked by white arrows. This stranding appears as linear areas of increased density within the fat surrounding the glands, suggesting inflammation and edema.\n\nThe image also provides evidence of the involvement of the superficial cervical fascia and platysma muscles, which can be inferred from the context. These structures may appear thickened due to the inflammatory process.\n\nThe bilateral involvement of the parotid glands is a common finding in viral sialadenitis, as it occurs in up to 75% of patients. However, it is important to note that a small minority of patients may present with only unilateral involvement.\n\nOverall, the image demonstrates the classic findings of viral sialadenitis, including bilateral enlargement of the salivary glands, fat stranding, and potential thickening of the superficial cervical fascia and platysma muscles. The patient's blood work, which tested positive for the parainfluenza virus, further supports the diagnosis.", "fig_caption": "Viral Sialadenitis. Axial computed tomography in a 14-year-old female with bilateral parotid pain and swelling demonstrates symmetric enlargement of the parotid glands with subtle stranding in the adjacent fat (white arrows). Blood work was positive for parainfluenza virus.", "in_text_mention": [{"tokens": "On imaging , classic findings include bilateral enlargement of the salivary glands , fat stranding , and thickening of the superficial cervical fascia and platysma muscles [ Figure 9 ] . [ 1213 ] Although bilateral involvement is seen in up to 75 % of patients , a small minority of patients may present with only unilateral involvement .", "entities": [{"type": "figure", "entity_id": "9", "start_tok": 27, "end_tok": 29}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 9"}
-{"question_id": 177, "image": "30910324_fig3.jpg", "pair_id": "30910324_fig3", "text": "Share a comprehensive rundown of the presented image\n", "gpt4_answer": "The image presents motion-corrected MRI data of a fetus with a double aortic arch at 32 weeks. It displays various anatomical structures, including the descending aorta, arterial duct, and left and right aortic arches, which are labeled accordingly. The image is divided into multiple panels, with the middle panel showing a contrast-enhanced MRI of the right-sided arch at 2 months postnatal age. \n\nIn the image, a ligamentous remnant of the left arch is predicted, marked by an asterisk. This finding is later confirmed during surgery, as shown in the right panel. Additionally, the distal remnant of the arterial duct is visible, which is analogous to the diverticulum of Kommerell and labeled with the letter \"K.\" The image is accompanied by a video that provides more detail on the subject.", "fig_caption": "Motion-corrected MRI data from a fetus with double aortic arch at 32 weeks Shown are the descending aorta (DAo), arterial duct (D), and left (L) and right (R) aortic arches. At 2 months postnatal age, contrast-enhanced MRI could show a right-sided arch (middle panel); however, a ligamentous remnant of the left arch was predicted on the basis of the fetal MRI findings (asterisk); this finding was confirmed at surgery (right panel). The distal remnant of the arterial duct\u2014analogous to the diverticulum of Kommerell\u2014is also seen (K). See video 3 for more detail.", "in_text_mention": [{"tokens": "Example images from 3D reconstructed data are shown in Figure 3 , Figure 4 , Figure 5 ; appendix p 3 ; videos 2-4 ) .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 9, "end_tok": 11}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 3"}
-{"question_id": 178, "image": "30911703_fig1.jpg", "pair_id": "30911703_fig1", "text": "Give an elaborate explanation of the image you see\n", "gpt4_answer": "The image is a computed tomography (CT) scan of the right lower extremity, focusing on the right peroneal artery. The CT scan uses contrast to enhance the visualization of the blood vessels in the area. In the image, you can see the right peroneal artery, which is a branch of the tibioperoneal trunk, a major blood vessel in the lower leg.\n\nThe main feature of the image is a saccular aneurysm located just distal to the tibioperoneal trunk. The aneurysm appears as a bulging, balloon-like structure in the right peroneal artery. The dimensions of the aneurysm are notable, as it is quite large in size. The shape of the aneurysm is irregular, and it is evident that the arterial wall has weakened and expanded in this area.\n\nThe surrounding structures in the image, such as bones, muscles, and other blood vessels, provide context for the location of the aneurysm. The CT scan offers a detailed view of the lower leg's anatomy, allowing for a clear understanding of the aneurysm's position and its relationship to nearby structures.\n\nOverall, the image provides a comprehensive view of the right peroneal artery aneurysm and its location in the lower leg, highlighting the importance of medical imaging in diagnosing and understanding vascular conditions.", "fig_caption": "Computed tomography scan of the right peroneal artery aneurysm.", "in_text_mention": [{"tokens": "A computed tomography scan with contrast of the right lower extremity demonstrated a 2.7- \u00d7 3.2- \u00d7 3.2-cm saccular aneurysm of his right peroneal artery just distal to the tibioperoneal trunk ( Fig 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 32, "end_tok": 34}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Fig\u00a01"}
-{"question_id": 179, "image": "30984437_fig2.jpg", "pair_id": "30984437_fig2", "text": "Clarify the contents of the displayed image with great detail\n", "gpt4_answer": "The image displays a chest computed tomography (CT) scan, which is a type of medical imaging technique used to visualize the internal structures of the chest. In this particular image, the focus is on the lungs.\n\nBoth lungs are visible in the image, and there are noticeable abnormalities present in the form of bilateral airspace consolidation and ground-glass opacity. Bilateral airspace consolidation refers to the presence of dense, opaque areas in both lungs, which can be seen as irregular, solid regions within the lung tissue. These areas of consolidation indicate that the airspaces within the lungs are filled with fluid, pus, or other cellular debris, which can be a sign of infection, inflammation, or other lung conditions.\n\nIn addition to the areas of consolidation, the image also shows ground-glass opacity, which is a term used to describe a hazy, semi-transparent appearance in the lung tissue. This opacity is less dense than the consolidation areas and can be seen as a lighter, more diffuse pattern throughout the lungs. Ground-glass opacity can be indicative of various lung conditions, including infections, interstitial lung diseases, or even early-stage lung cancer.\n\nOverall, the image provides a detailed view of the lung abnormalities, highlighting the bilateral airspace consolidation and ground-glass opacity, which can be helpful in diagnosing and managing various lung conditions.", "fig_caption": "Chest computed tomography revealed bilateral airspace consolidation and ground-glass opacity.", "in_text_mention": [{"tokens": "Chest computed tomography ( CT ) revealed bilateral airspace consolidation and ground-glass opacity ( Figure 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 14, "end_tok": 16}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 2"}
-{"question_id": 180, "image": "31404134_F5.jpg", "pair_id": "31404134_F5", "text": "Analyze the image in a comprehensive and detailed manner\n", "gpt4_answer": "The image is a chest x-ray taken a few weeks after the patient started hemodialysis. In the image, the patient's chest cavity can be seen, including the lungs, heart, and ribcage. The overall appearance of the lungs is clear, indicating that there is no pleural effusion present. This suggests that the patient's condition has improved since starting hemodialysis.\n\nAdditionally, a perm catheter can be observed in the image. The catheter is positioned correctly, allowing for efficient hemodialysis treatment. The placement of the catheter is important for the patient's ongoing care and treatment.\n\nOverall, the chest x-ray image demonstrates the patient's progress after starting hemodialysis, with no signs of pleural effusion and a properly placed perm catheter.", "fig_caption": "Chest x-ray, few weeks after starting hemodialysis, showed no pleural effusion. Perm catheter is in place", "in_text_mention": [{"tokens": "Follow-up evaluation showed neither recurrence of symptoms nor recurrence of pleural effusion on chest x-ray ( see Figure 5 ) .", "entities": [{"type": "figure", "entity_id": "5", "start_tok": 17, "end_tok": 19}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 5"}
-{"question_id": 181, "image": "32009718_F1.jpg", "pair_id": "32009718_F1", "text": "Describe the following image in detail\n", "gpt4_answer": "The image is a chest X-ray of a patient, showing a detailed view of their lungs. In the right lung, there are multiple patchy areas of consolidation distributed throughout the lung fields. These areas appear as irregular, dense regions, indicating possible infection or inflammation. Similarly, in the left lung, the upper zone also displays patchy consolidation, suggesting a similar issue. The overall appearance of the lungs in the image highlights the presence of a medical condition that affects both lungs, with a more pronounced impact on the right side.", "fig_caption": "Chest X-ray of a patient at presentation. Patchy areas of consolidation seen throughout the right lung and left upper zone", "in_text_mention": [{"tokens": "Chest roentgenogram showed patchy areas of consolidation throughout the right lung fields and left upper lobe [ Figure 1 ] .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 17, "end_tok": 19}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 1"}
-{"question_id": 182, "image": "32418875_fig1.jpg", "pair_id": "32418875_fig1", "text": "Share a comprehensive rundown of the presented image\n", "gpt4_answer": "The image presents a flowchart illustrating the adult pathway to screen for acute heart failure in COVID-19 patients admitted to the hospital. The flowchart begins with the patient's admission to the emergency department. From there, several tests are conducted, including electrolytes, urea and creatine tests, liver function tests, C-reactive protein tests, lactate dehydrogenase tests, chest X-rays, and electrocardiographs.\n\nBased on the results of these tests, the flowchart guides the healthcare professionals through various decision points to determine the appropriate course of action. This may involve admitting the patient to a high dependency unit or an intensive care unit, depending on the severity of their condition. In some cases, a transthoracic echocardiogram may be performed to assess left ventricular function.\n\nThe flowchart aims to help identify COVID-19 patients at risk for severe heart failure and circulatory collapse while balancing the need to protect healthcare workers from virus exposure and preserving personal protective equipment.", "fig_caption": "Adult pathway to screen for acute heart failure in COVID-19 patients admitted to hospital. Abbreviations: ED, emergency department; EUC, electrolytes, urea and creatine; LFT, liver function test; CRP, C-reactive protein; LDH, lactate dehydrogenase; CXR, chest X-ray; ECG, electrocardiograph; HDU, high dependency unit; ICU, intensive care unit; TTE, transthoracic echocardiogram; LV, left ventricular.", "in_text_mention": [{"tokens": "We suggest the following algorithm ( Figure 1 ) , to better identify COVID-19 patients at risk for severe heart failure and circulatory collapse , whilst balancing the need to protect health care workers from virus exposure and to preserve personal protective equipment ( PPE ) :", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 6, "end_tok": 8}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure\u00a01"}
-{"question_id": 183, "image": "32548036_F5.jpg", "pair_id": "32548036_F5", "text": "Examine the image closely and share its details\n", "gpt4_answer": "The image displays a gross specimen, which is a sample of biological tissue taken for examination. In this case, the specimen appears to have been removed from a patient during a surgical procedure. The tissue sample is shown in its natural state, without any staining or magnification.\n\nThe specimen is placed on a clean, flat surface, likely a tray or a petri dish, to provide a clear view of its features. The tissue has a distinct color, texture, and shape, which can provide valuable information about the patient's condition. The size of the specimen can also be observed, giving an idea of the extent of the affected area.\n\nThe image may also show some tools or instruments used during the procedure, such as forceps or a scalpel, which can help to provide context for the scale of the specimen. Additionally, there might be labels or annotations on the image to indicate specific areas of interest or to provide further information about the sample.\n\nOverall, the image serves as a visual representation of the gross specimen, allowing researchers and medical professionals to study its features and gain insights into the patient's condition.", "fig_caption": "Gross specimen.", "in_text_mention": null, "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "detailed_description", "fig_label": "Figure 5"}
-{"question_id": 184, "image": "32715172_fig2.jpg", "pair_id": "32715172_fig2", "text": "Narrate the contents of the image with precision\n", "gpt4_answer": "The image displays a preoperative aortography, which is a medical imaging technique used to visualize the anatomy of the aorta. In this particular image, the aorta is shown to be consistent with the findings from a computed tomography (CT) scan. The aortography reveals the structure and any potential abnormalities in the aorta, allowing for a clear understanding of the patient's anatomy.\n\nThe image likely shows the aorta in detail, with its various branches and blood vessels visible. The contrast between the aorta and the surrounding tissues is evident, making it easy to identify the specific anatomical features. The image also highlights the suitability of the patient's anatomy for the proposed repair, as confirmed by the intraoperative flush abdominal aortography. Overall, the image provides a clear and informative visualization of the aorta, which is crucial for planning and executing the necessary medical procedures.", "fig_caption": "Preoperative aortography demonstrating anatomy consistent with the computed tomography scan.", "in_text_mention": [{"tokens": "Intraoperative flush abdominal aortography confirmed computed tomography angiography findings and a suitable anatomy for the proposed repair ( Fig 2 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 18, "end_tok": 20}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Fig\u00a02"}
-{"question_id": 185, "image": "32867401_materials-13-03795-f015.jpg", "pair_id": "32867401_materials-13-03795-f015", "text": "Break down the elements of the image in a detailed manner\n", "gpt4_answer": "The image is a schematic representation of a multi-modal Janus nanoparticulate system. This system consists of several components that contribute to its functionality in biomedical applications.\n\nAt the center of the image, there is a core/shell face composed of superparamagnetic iron oxide nanoparticles (SPION) and mesoporous silica nanoparticles (MSN). This core/shell structure is responsible for the T2 magnetic resonance imaging (MRI) contrast capabilities of the system.\n\nOn one side of the core/shell face, there is a gold nanoparticle face. This gold interface is responsible for providing the possibility of computed tomography (CT) imaging.\n\nThe gold nanoparticle face is also modified with a fluorescent dye, which is depicted in the image as a distinct element attached to the gold interface. This dye, Alexa Fluor\u00ae 647, allows for optical imaging of the system.\n\nOn the other side, the MSN face is functionalized with a tumor-targeting peptide, cRDG. This peptide is shown in the schematic as a separate element attached to the MSN face, enabling active tumor targeting.\n\nOverall, the image provides a clear and detailed visualization of the multi-modal Janus nanoparticulate system, highlighting its various components and their roles in imaging and tumor targeting.", "fig_caption": "A schematic representing a multi-modal Janus nanoparticulate system composed of a SPION/MSN core/shell face and Au nanoparticle face. T2 MRI contrast capabilities arise from the encapsulated SPION with Au nanoparticles providing the possibility for CT imaging. As shown in this schematic, the system is functionalised with a fluorescent dye (Alexa Fluor\u00ae 647) for optical imaging in addition to cRDG for active tumour targeting. Reprinted with permission from reference [178]. Copyright (2018) American Chemical Society.", "in_text_mention": [{"tokens": "The gold interface was modified with a fluorescent dye ( Alexa Fluor\u00ae 647 ) and the MSN face with a tumour targeting peptide cRDG ( Figure 15 ) .", "entities": [{"type": "figure", "entity_id": "15", "start_tok": 25, "end_tok": 27}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 15"}
-{"question_id": 186, "image": "32952957_F4.jpg", "pair_id": "32952957_F4", "text": "Clarify the contents of the displayed image with great detail\n", "gpt4_answer": "The image consists of two parts. In the top section, there are representative microphotographs of histological slices from the left ventricles of rat hearts. These slices are stained with Sirius Red, which highlights collagen fibers in red. The microphotographs display heart tissue samples from both short-term T3 treated rats and vehicle-treated rats, allowing for comparison between the two groups. A scale bar of 25 \u00b5m is provided to indicate the size of the structures in the images.\n\nIn the bottom section of the image, there is a quantitative analysis of the collagen-positive areas in the left ventricles of the different animal groups. This analysis is presented as a bar graph, with the percentage of positively stained collagen area expressed as a percentage of the total area. The graph includes data for short-term hyperthyroidism induced by daily subcutaneous injections of T3 for 1, 3, and 10 days (T31d, T33d, and T310d), as well as control animals injected daily with the vehicle for 1, 3, and 10 days (V1d, V3d, and V10d). The values are represented as means\u00b1SEM, and the statistical analysis (one-way ANOVA) indicates no significant differences between the groups.", "fig_caption": "Absence of signs of fibrosis in the left ventricles of 3, 5, 3\u2019 triiodo-L-thyronine treated rats. (top) Representative microphotographs of histological slices stained with Sirius Red of hearts from short-term T3 treated and vehicle-treated rats (red color indicates collagen fibers; scale bar 25 \u00b5m). (bottom) Quantitative analysis of left ventricular collagen-positive areas in each of the different animal groups. The positively stained collagen area was expressed as a percentage of the total area. Short-term hyperthyroidism was induced by the daily subcutaneous injection of T3 (500 \u03bcg.kg-1) for 1 (T31d), 3 (T33d), and 10 (T310d) days. Control animals were injected daily with the vehicle for 1 (V1d), 3 (V3d), and 10 (V10d) days. Values are means\u00b1SEM. P not significant; n= 60 randomly chosen frames from 6 hearts per group (one-way ANOVA)", "in_text_mention": [{"tokens": "Sirius Red staining to determine collagen deposition showed no more connective tissue in the left ventricles of T31d , T33d , and T310d rats compared with their respective temporal controls ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 31, "end_tok": 33}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "detailed_description", "fig_label": "Figure 4"}
-{"question_id": 187, "image": "33708306_f0004.jpg", "pair_id": "33708306_f0004", "text": "Illustrate the image through a descriptive explanation\n", "gpt4_answer": "The image is a chest X-ray focusing on the mediastinal region. In this particular X-ray, there is a noticeable widening of the mediastinum, which is an abnormal finding. To help viewers identify this widening, a white arrow is pointing directly at the area of concern. The overall image provides a clear visualization of the chest structures, including the lungs and heart, allowing for a better understanding of the patient's condition.", "fig_caption": "Chest X ray - case 2: mediastinal widening (white arrow)", "in_text_mention": [{"tokens": "Chest X ray showed a mediastinal widening ( Figure 4 ) .", "entities": [{"type": "figure", "entity_id": "4", "start_tok": 8, "end_tok": 10}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 4"}
-{"question_id": 188, "image": "33864678_f1-amjcaserep-22-e930902.jpg", "pair_id": "33864678_f1-amjcaserep-22-e930902", "text": "Write an exhaustive depiction of the given image\n", "gpt4_answer": "The image is a computed tomography (CT) scan displaying a cross-sectional view of a patient's thoracic region. The focus of the image is on the mass that arises from the posterior costal arc of the eighth rib on the patient's left side. The mass appears to be well-defined and localized, with clear boundaries separating it from the surrounding tissues.\n\nThe eighth rib is visible in the image, and the mass can be seen originating from its posterior costal arc. The rib's structure is also discernible, with the curvature of the rib and its connection to the spine clearly visible. The patient's left side is marked for orientation purposes.\n\nThe CT scan also shows the medulla, which is not involved with the mass. The medulla appears to be intact and separate from the mass, indicating that the mass has not infiltrated or affected this area. The surrounding tissues and organs, such as the lungs and the diaphragm, can also be seen in the image, providing context for the location of the mass.\n\nOverall, the image provides a detailed view of the mass and its relationship to the eighth rib and the medulla, allowing for a better understanding of the patient's condition.", "fig_caption": "A computed tomography scan shows that the mass arises from the posterior costal arc of the eighth rib on the patient\u2019s left side and does not involve the medulla.", "in_text_mention": null, "domain": {"chest_xray": false, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 1."}
-{"question_id": 189, "image": "33935946_F1.jpg", "pair_id": "33935946_F1", "text": "Describe the following image in detail\n", "gpt4_answer": "The image is a selected slice from a multi-modal MRI dataset of a patient with Parkinson's disease. It is a high-resolution, grayscale image that showcases the intricate details of the patient's brain anatomy. The MRI scan was performed using a 3T Siemens Skyra MR scanner at a university medical center.\n\nIn the image, various brain structures can be observed, such as the cerebral cortex, white matter, and deep brain nuclei. The contrast between different tissue types is evident, allowing for a clear distinction between gray and white matter. The image also highlights any potential abnormalities or changes in the brain structure that may be associated with Parkinson's disease.\n\nOverall, the image provides valuable insights into the brain's anatomy and the potential effects of Parkinson's disease on the patient's brain.", "fig_caption": "Selected slice from a multi-modal MRI dataset of a patient with Parkinson's disease.", "in_text_mention": [{"tokens": "All participants were scanned at the University Medical Center Hamburg-Eppendorf , Germany , using a 3T Siemens Skyra MR scanner ( Figure 1 ) .", "entities": [{"type": "figure", "entity_id": "1", "start_tok": 21, "end_tok": 23}]}], "domain": {"chest_xray": false, "mri": true, "ct_scan": false, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 1"}
-{"question_id": 190, "image": "34026556_fig2.jpg", "pair_id": "34026556_fig2", "text": "Describe the following image in detail\n", "gpt4_answer": "The image displays a gross anatomical view of the appendix and right kidney. The appendix appears to be abnormally adhered to the right kidney, indicating a pathological condition. The two organs are visibly connected, possibly due to the formation of a fistulous tract. The image likely shows the organs after they have been surgically removed from the patient during an appendectomy. The overall appearance of the appendix and right kidney may exhibit signs of inflammation or other abnormalities related to the medical condition.", "fig_caption": "Gross image of appendix adhered to right kidney.", "in_text_mention": [{"tokens": "An appendectomy was performed to allow for complete removal of the fistulous tract along with the right kidney ( Fig. 2 , Fig. 3 ) .", "entities": [{"type": "figure", "entity_id": "2", "start_tok": 19, "end_tok": 21}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": false, "gross": true}, "type": "detailed_description", "fig_label": "Fig. 2"}
-{"question_id": 191, "image": "34267836_F7.jpg", "pair_id": "34267836_F7", "text": "Examine the image closely and share its details\n", "gpt4_answer": "The image displays a microscopic view of a metastatic liver lesion at 400x magnification, using hematoxylin and eosin (H&E) staining. The stained tissue sample reveals the presence of mixed adenocarcinoma and high-grade neuroendocrine carcinoma. The adenocarcinoma cells appear as glandular structures, while the neuroendocrine carcinoma cells exhibit a more dispersed pattern. The H&E staining highlights the cellular details, allowing for the differentiation between the two types of cancer cells. The image provides a clear visualization of the coexistence of these two distinct cell types within the metastatic liver lesion.", "fig_caption": "Mixed adenocarcinoma high-grade neuroendocrine carcinoma in metastatic liver lesion (H&E, original \u00d7 400). H&E: hematoxylin and eosin stain.", "in_text_mention": [{"tokens": "Liver biopsy also showed mixed adenocarcinoma and neuroendocrine carcinoma , high-grade ( Fig. 7 ) .", "entities": [{"type": "figure", "entity_id": "7", "start_tok": 12, "end_tok": 14}]}], "domain": {"chest_xray": false, "mri": false, "ct_scan": false, "histology": true, "gross": false}, "type": "detailed_description", "fig_label": "Figure 7"}
-{"question_id": 192, "image": "34574873_healthcare-09-01099-f003.jpg", "pair_id": "34574873_healthcare-09-01099-f003", "text": "Provide a detailed description of the given image\n", "gpt4_answer": "The image displays a collection of representative sample images of chest X-rays and CT scans. These images are arranged in a grid-like format, showcasing the variety of cases included in the mixed dataset used for analysis. The chest X-rays exhibit different lung conditions, with varying degrees of clarity and contrast, while the CT scans provide cross-sectional views of the chest, highlighting the internal structures and any abnormalities present. The image serves as a visual representation of the diverse dataset used in the study, emphasizing the range of cases and imaging techniques employed in the analysis.", "fig_caption": "Representative sample images of chest X-rays and CT scans used in the mixed dataset adopted for analysis.", "in_text_mention": [{"tokens": "Figure 3 presents a set of representative images used in the analysis .", "entities": [{"type": "figure", "entity_id": "3", "start_tok": 0, "end_tok": 2}]}], "domain": {"chest_xray": true, "mri": false, "ct_scan": true, "histology": false, "gross": false}, "type": "detailed_description", "fig_label": "Figure 3"}
diff --git a/LLaVA-Med/docs/llava_med_performance.md b/LLaVA-Med/docs/llava_med_performance.md
deleted file mode 100644
index 55028b366fc2703761632a7108d5f3c5153d6101..0000000000000000000000000000000000000000
--- a/LLaVA-Med/docs/llava_med_performance.md
+++ /dev/null
@@ -1,31 +0,0 @@
-## LLaVA-Med-1.5 Performance
-
-
- 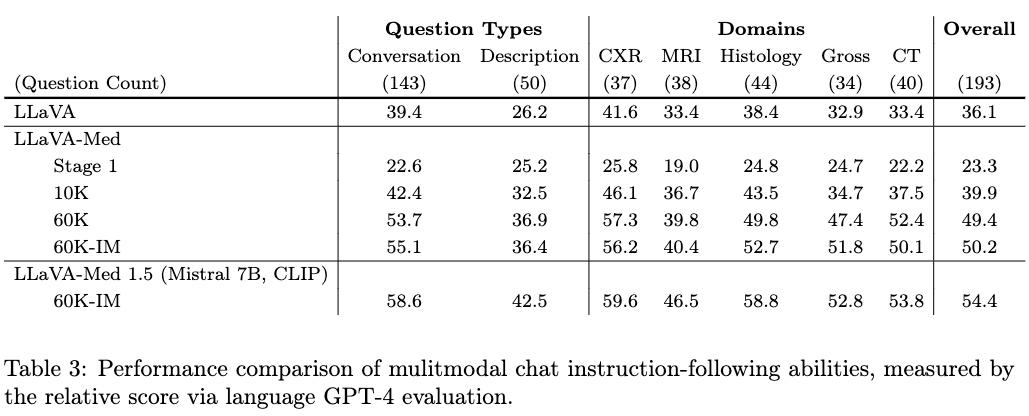
-
- *Performance comparison of mulitmodal chat instruction-following abilities, measured by the relative score via language GPT-4 evaluation.*
-
-
-
-## LLaVA-Med-1.0 Performance
-
-
- 
-
- *Example 1: comparison of medical visual chat. The language-only GPT-4 is considered as the performance upper bound, as the golden captions and inline mentions are fed into GPT-4 as the context, without requiring the model to understand the raw image.*
-
-
-
- 
-
- *Example 2: comparison of medical visual chat. LLaVA tends to halluciate or refuse to provide domain-specific knowledgable response.*
-
-
-
-
- 
-
- *Performance comparison of fine-tuned LLaVA-Med on established Medical QVA datasets.*
-
-
-
diff --git a/LLaVA-Med/download_data.sh b/LLaVA-Med/download_data.sh
deleted file mode 100644
index c38b9bb94b2766d39e23f57ee4a4521126851442..0000000000000000000000000000000000000000
--- a/LLaVA-Med/download_data.sh
+++ /dev/null
@@ -1,35 +0,0 @@
-#!/bin/bash
-
-mkdir data/alignment
-cd data/alignment
-
-wget https://hanoverprod.z21.web.core.windows.net/med_llava/alignment/llava_med_alignment_500k.json
-
-cd ..
-
-mkdir instruct
-cd instruct
-
-wget https://hanoverprod.z21.web.core.windows.net/med_llava/instruct/llava_med_instruct_10k.json
-wget https://hanoverprod.z21.web.core.windows.net/med_llava/instruct/llava_med_instruct_60k.json
-wget https://hanoverprod.z21.web.core.windows.net/med_llava/instruct/llava_med_instruct_60k_inline_mention.json
-wget https://hanoverprod.z21.web.core.windows.net/med_llava/instruct/llava_med_instruct_fig_captions.json
-cd ..
-
-mkdir eval
-cd eval
-
-wget https://hanoverprod.z21.web.core.windows.net/med_llava/eval/llava_med_eval_qa50_qa.jsonl
-wget https://hanoverprod.z21.web.core.windows.net/med_llava/eval/llava_med_eval_qa50_fig_captions.json
-wget https://hanoverprod.z21.web.core.windows.net/med_llava/eval/llava_med_qa50_instruct_caption_in_text_cleaned-60k-3epoch.json
-
-cd ..
-
-wget https://hanoverprod.z21.web.core.windows.net/med_llava/llava_med_image_urls.jsonl
-mkdir pmc_articles
-mkdir images
-
-cd ..
-
-pip install tqdm
-python llava/data/download_images.py --input_path data/llava_med_image_urls.jsonl --pmc_output_path data/pmc_articles/ --images_output_path data/images
\ No newline at end of file
diff --git a/LLaVA-Med/images/llava_logo.png b/LLaVA-Med/images/llava_logo.png
deleted file mode 100644
index 567428adb29c03dd83c1f08be6b4e972af453630..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/images/llava_logo.png and /dev/null differ
diff --git a/LLaVA-Med/images/llava_med_chat.png b/LLaVA-Med/images/llava_med_chat.png
deleted file mode 100644
index eb9573ed246a3aa84f8981da285d402a79a1ea56..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/images/llava_med_chat.png and /dev/null differ
diff --git a/LLaVA-Med/images/llava_med_chat_example1.png b/LLaVA-Med/images/llava_med_chat_example1.png
deleted file mode 100644
index 7677c1428e9c788ed9a3e3baf89f5d73967bbade..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/images/llava_med_chat_example1.png and /dev/null differ
diff --git a/LLaVA-Med/images/llava_med_chat_example2.png b/LLaVA-Med/images/llava_med_chat_example2.png
deleted file mode 100644
index 5c9afd6942e46ab975e680bb48ccebaffcfbd8ca..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/images/llava_med_chat_example2.png and /dev/null differ
diff --git a/LLaVA-Med/images/llava_med_dataset.png b/LLaVA-Med/images/llava_med_dataset.png
deleted file mode 100644
index f4f3cca17c3a4fa2126bc2eebef4fd5ff215a658..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/images/llava_med_dataset.png and /dev/null differ
diff --git a/LLaVA-Med/images/llava_med_logo.png b/LLaVA-Med/images/llava_med_logo.png
deleted file mode 100644
index 7b1224978ff229aff77808577054d0ae4247572b..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/images/llava_med_logo.png and /dev/null differ
diff --git a/LLaVA-Med/images/llava_med_pipeline.png b/LLaVA-Med/images/llava_med_pipeline.png
deleted file mode 100644
index 1f3ab4017b780f5c74517b5b4a6c05f434b99750..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/images/llava_med_pipeline.png and /dev/null differ
diff --git a/LLaVA-Med/images/llava_med_vqa.png b/LLaVA-Med/images/llava_med_vqa.png
deleted file mode 100644
index cd695741d8408cb050de7a02b307793b64148ca2..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/images/llava_med_vqa.png and /dev/null differ
diff --git a/LLaVA-Med/llava/__init__.py b/LLaVA-Med/llava/__init__.py
deleted file mode 100644
index e69de29bb2d1d6434b8b29ae775ad8c2e48c5391..0000000000000000000000000000000000000000
diff --git a/LLaVA-Med/llava/constants.py b/LLaVA-Med/llava/constants.py
deleted file mode 100644
index 374be090510b302de9882d880c755787a8eafe11..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/constants.py
+++ /dev/null
@@ -1,13 +0,0 @@
-CONTROLLER_HEART_BEAT_EXPIRATION = 30
-WORKER_HEART_BEAT_INTERVAL = 15
-
-LOGDIR = "."
-
-# Model Constants
-IGNORE_INDEX = -100
-IMAGE_TOKEN_INDEX = -200
-DEFAULT_IMAGE_TOKEN = ""
-DEFAULT_IMAGE_PATCH_TOKEN = ""
-DEFAULT_IM_START_TOKEN = ""
-DEFAULT_IM_END_TOKEN = ""
-IMAGE_PLACEHOLDER = ""
diff --git a/LLaVA-Med/llava/conversation.py b/LLaVA-Med/llava/conversation.py
deleted file mode 100644
index 92550d7e789cd0cde19e137f733ca6b7ed2adbac..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/conversation.py
+++ /dev/null
@@ -1,439 +0,0 @@
-import dataclasses
-from enum import auto, Enum
-from typing import List, Tuple
-import base64
-from io import BytesIO
-from PIL import Image
-
-
-class SeparatorStyle(Enum):
- """Different separator style."""
- SINGLE = auto()
- TWO = auto()
- MPT = auto()
- PLAIN = auto()
- LLAMA_2 = auto()
- MISTRAL = auto()
-
-
-@dataclasses.dataclass
-class Conversation:
- """A class that keeps all conversation history."""
- system: str
- roles: List[str]
- messages: List[List[str]]
- offset: int
- sep_style: SeparatorStyle = SeparatorStyle.SINGLE
- sep: str = "###"
- sep2: str = None
- version: str = "Unknown"
-
- skip_next: bool = False
-
- def get_prompt(self):
- messages = self.messages
- if len(messages) > 0 and type(messages[0][1]) is tuple:
- messages = self.messages.copy()
- init_role, init_msg = messages[0].copy()
- init_msg = init_msg[0].replace("", "").strip()
- if 'mmtag' in self.version:
- messages[0] = (init_role, init_msg)
- messages.insert(0, (self.roles[0], ""))
- messages.insert(1, (self.roles[1], "Received."))
- else:
- messages[0] = (init_role, "\n" + init_msg)
-
- if self.sep_style == SeparatorStyle.SINGLE:
- ret = self.system + self.sep
- for role, message in messages:
- if message:
- if type(message) is tuple:
- message, _, _ = message
- ret += role + ": " + message + self.sep
- else:
- ret += role + ":"
- elif self.sep_style == SeparatorStyle.TWO:
- seps = [self.sep, self.sep2]
- ret = self.system + seps[0]
- for i, (role, message) in enumerate(messages):
- if message:
- if type(message) is tuple:
- message, _, _ = message
- sep = seps[i % 2]
- sep = "{0} ".format(self.sep2) if sep == self.sep2 else self.sep
- ret += role + ": " + message.strip() + sep
- else:
- ret += role + ":"
- ret = ret.strip()
- elif self.sep_style == SeparatorStyle.MPT:
- ret = self.system + self.sep
- for role, message in messages:
- if message:
- if type(message) is tuple:
- message, _, _ = message
- ret += role + message + self.sep
- else:
- ret += role
- elif self.sep_style == SeparatorStyle.LLAMA_2:
- wrap_sys = lambda msg: f"<>\n{msg}\n<>\n\n" if len(msg) > 0 else msg
- wrap_inst = lambda msg: f"[INST] {msg} [/INST]"
- ret = ""
-
- for i, (role, message) in enumerate(messages):
- if i == 0:
- assert message, "first message should not be none"
- assert role == self.roles[0], "first message should come from user"
- if message:
- if type(message) is tuple:
- message, _, _ = message
- if i == 0: message = wrap_sys(self.system) + message
- if i % 2 == 0:
- message = wrap_inst(message)
- ret += self.sep + message
- else:
- ret += " " + message + " " + self.sep2
- else:
- ret += ""
- ret = ret.lstrip(self.sep)
- elif self.sep_style == SeparatorStyle.PLAIN:
- seps = [self.sep, self.sep2]
- ret = self.system
- for i, (role, message) in enumerate(messages):
- if message:
- if type(message) is tuple:
- message, _, _ = message
- ret += message + seps[i % 2]
- else:
- ret += ""
- elif self.sep_style == SeparatorStyle.MISTRAL:
- # reference: https://docs.mistral.ai/models/
- wrap_sys = lambda msg: f"{msg}"
- wrap_inst = lambda msg: f"[INST] {msg} [/INST]"
- ret = ""
- for i, (role, message) in enumerate(messages):
- if i == 0:
- assert message, "first message should not be none"
- assert role == self.roles[0], "first message should come from user"
- if message:
- if type(message) is tuple:
- message, _, _ = message
- if i == 0: message = self.system + " " + message.strip()
- if i % 2 == 0:
- message = wrap_inst(message)
- ret += message
- else:
- ret += wrap_sys(message)
- else:
- ret += ""
- # wrap_sys = lambda msg: f"\n{msg}\n\n"
- # wrap_inst = lambda msg: f"[INST] {msg} [/INST]"
- # ret = ""
- # for i, (role, message) in enumerate(messages):
- # if i == 0:
- # assert message, "first message should not be none"
- # assert role == self.roles[0], "first message should come from user"
- # if message:
- # if type(message) is tuple:
- # message, _, _ = message
- # if i == 0: message = wrap_sys(self.system) + message
- # if i % 2 == 0:
- # message = wrap_inst(message)
- # ret += message if i != 0 else self.sep + message
- # else:
- # # NOTE-JW: we need to add " " to strictly follow Mistral Instruction Format
- # ret += " " + message + " " + self.sep2
- # # ret += " " + wrap_sys(message)
- # else:
- # ret += ""
- else:
- raise ValueError(f"Invalid style: {self.sep_style}")
-
- return ret
-
- def append_message(self, role, message):
- self.messages.append([role, message])
-
- def get_images(self, return_pil=False):
- images = []
- for i, (role, msg) in enumerate(self.messages[self.offset:]):
- if i % 2 == 0:
- if type(msg) is tuple:
- import base64
- from io import BytesIO
- from PIL import Image
- msg, image, image_process_mode = msg
- if image_process_mode == "Pad":
- def expand2square(pil_img, background_color=(122, 116, 104)):
- width, height = pil_img.size
- if width == height:
- return pil_img
- elif width > height:
- result = Image.new(pil_img.mode, (width, width), background_color)
- result.paste(pil_img, (0, (width - height) // 2))
- return result
- else:
- result = Image.new(pil_img.mode, (height, height), background_color)
- result.paste(pil_img, ((height - width) // 2, 0))
- return result
- image = expand2square(image)
- elif image_process_mode in ["Default", "Crop"]:
- pass
- elif image_process_mode == "Resize":
- image = image.resize((336, 336))
- else:
- raise ValueError(f"Invalid image_process_mode: {image_process_mode}")
- max_hw, min_hw = max(image.size), min(image.size)
- aspect_ratio = max_hw / min_hw
- max_len, min_len = 800, 400
- shortest_edge = int(min(max_len / aspect_ratio, min_len, min_hw))
- longest_edge = int(shortest_edge * aspect_ratio)
- W, H = image.size
- if longest_edge != max(image.size):
- if H > W:
- H, W = longest_edge, shortest_edge
- else:
- H, W = shortest_edge, longest_edge
- image = image.resize((W, H))
- if return_pil:
- images.append(image)
- else:
- buffered = BytesIO()
- image.save(buffered, format="PNG")
- img_b64_str = base64.b64encode(buffered.getvalue()).decode()
- images.append(img_b64_str)
- return images
-
- def to_gradio_chatbot(self):
- ret = []
- for i, (role, msg) in enumerate(self.messages[self.offset:]):
- if i % 2 == 0:
- if type(msg) is tuple:
- import base64
- from io import BytesIO
- msg, image, image_process_mode = msg
- max_hw, min_hw = max(image.size), min(image.size)
- aspect_ratio = max_hw / min_hw
- max_len, min_len = 800, 400
- shortest_edge = int(min(max_len / aspect_ratio, min_len, min_hw))
- longest_edge = int(shortest_edge * aspect_ratio)
- W, H = image.size
- if H > W:
- H, W = longest_edge, shortest_edge
- else:
- H, W = shortest_edge, longest_edge
- image = image.resize((W, H))
- buffered = BytesIO()
- image.save(buffered, format="JPEG")
- img_b64_str = base64.b64encode(buffered.getvalue()).decode()
- img_str = f' '
- msg = img_str + msg.replace('', '').strip()
- ret.append([msg, None])
- else:
- ret.append([msg, None])
- else:
- ret[-1][-1] = msg
- return ret
-
- def copy(self):
- return Conversation(
- system=self.system,
- roles=self.roles,
- messages=[[x, y] for x, y in self.messages],
- offset=self.offset,
- sep_style=self.sep_style,
- sep=self.sep,
- sep2=self.sep2,
- version=self.version)
-
- def dict(self):
- if len(self.get_images()) > 0:
- return {
- "system": self.system,
- "roles": self.roles,
- "messages": [[x, y[0] if type(y) is tuple else y] for x, y in self.messages],
- "offset": self.offset,
- "sep": self.sep,
- "sep2": self.sep2,
- }
- return {
- "system": self.system,
- "roles": self.roles,
- "messages": self.messages,
- "offset": self.offset,
- "sep": self.sep,
- "sep2": self.sep2,
- }
-
-
-conv_vicuna_v0 = Conversation(
- system="A chat between a curious human and an artificial intelligence assistant. "
- "The assistant gives helpful, detailed, and polite answers to the human's questions.",
- roles=("Human", "Assistant"),
- messages=(
- ("Human", "What are the key differences between renewable and non-renewable energy sources?"),
- ("Assistant",
- "Renewable energy sources are those that can be replenished naturally in a relatively "
- "short amount of time, such as solar, wind, hydro, geothermal, and biomass. "
- "Non-renewable energy sources, on the other hand, are finite and will eventually be "
- "depleted, such as coal, oil, and natural gas. Here are some key differences between "
- "renewable and non-renewable energy sources:\n"
- "1. Availability: Renewable energy sources are virtually inexhaustible, while non-renewable "
- "energy sources are finite and will eventually run out.\n"
- "2. Environmental impact: Renewable energy sources have a much lower environmental impact "
- "than non-renewable sources, which can lead to air and water pollution, greenhouse gas emissions, "
- "and other negative effects.\n"
- "3. Cost: Renewable energy sources can be more expensive to initially set up, but they typically "
- "have lower operational costs than non-renewable sources.\n"
- "4. Reliability: Renewable energy sources are often more reliable and can be used in more remote "
- "locations than non-renewable sources.\n"
- "5. Flexibility: Renewable energy sources are often more flexible and can be adapted to different "
- "situations and needs, while non-renewable sources are more rigid and inflexible.\n"
- "6. Sustainability: Renewable energy sources are more sustainable over the long term, while "
- "non-renewable sources are not, and their depletion can lead to economic and social instability.\n")
- ),
- offset=2,
- sep_style=SeparatorStyle.SINGLE,
- sep="###",
-)
-
-conv_vicuna_v1 = Conversation(
- system="A chat between a curious user and an artificial intelligence assistant. "
- "The assistant gives helpful, detailed, and polite answers to the user's questions.",
- roles=("USER", "ASSISTANT"),
- version="v1",
- messages=(),
- offset=0,
- sep_style=SeparatorStyle.TWO,
- sep=" ",
- sep2="",
-)
-
-conv_llama_2 = Conversation(
- system="""You are a helpful, respectful and honest assistant. Always answer as helpfully as possible, while being safe. Your answers should not include any harmful, unethical, racist, sexist, toxic, dangerous, or illegal content. Please ensure that your responses are socially unbiased and positive in nature.
-
-If a question does not make any sense, or is not factually coherent, explain why instead of answering something not correct. If you don't know the answer to a question, please don't share false information.""",
- roles=("USER", "ASSISTANT"),
- version="llama_v2",
- messages=(),
- offset=0,
- sep_style=SeparatorStyle.LLAMA_2,
- sep="
'
- msg = img_str + msg.replace('', '').strip()
- ret.append([msg, None])
- else:
- ret.append([msg, None])
- else:
- ret[-1][-1] = msg
- return ret
-
- def copy(self):
- return Conversation(
- system=self.system,
- roles=self.roles,
- messages=[[x, y] for x, y in self.messages],
- offset=self.offset,
- sep_style=self.sep_style,
- sep=self.sep,
- sep2=self.sep2,
- version=self.version)
-
- def dict(self):
- if len(self.get_images()) > 0:
- return {
- "system": self.system,
- "roles": self.roles,
- "messages": [[x, y[0] if type(y) is tuple else y] for x, y in self.messages],
- "offset": self.offset,
- "sep": self.sep,
- "sep2": self.sep2,
- }
- return {
- "system": self.system,
- "roles": self.roles,
- "messages": self.messages,
- "offset": self.offset,
- "sep": self.sep,
- "sep2": self.sep2,
- }
-
-
-conv_vicuna_v0 = Conversation(
- system="A chat between a curious human and an artificial intelligence assistant. "
- "The assistant gives helpful, detailed, and polite answers to the human's questions.",
- roles=("Human", "Assistant"),
- messages=(
- ("Human", "What are the key differences between renewable and non-renewable energy sources?"),
- ("Assistant",
- "Renewable energy sources are those that can be replenished naturally in a relatively "
- "short amount of time, such as solar, wind, hydro, geothermal, and biomass. "
- "Non-renewable energy sources, on the other hand, are finite and will eventually be "
- "depleted, such as coal, oil, and natural gas. Here are some key differences between "
- "renewable and non-renewable energy sources:\n"
- "1. Availability: Renewable energy sources are virtually inexhaustible, while non-renewable "
- "energy sources are finite and will eventually run out.\n"
- "2. Environmental impact: Renewable energy sources have a much lower environmental impact "
- "than non-renewable sources, which can lead to air and water pollution, greenhouse gas emissions, "
- "and other negative effects.\n"
- "3. Cost: Renewable energy sources can be more expensive to initially set up, but they typically "
- "have lower operational costs than non-renewable sources.\n"
- "4. Reliability: Renewable energy sources are often more reliable and can be used in more remote "
- "locations than non-renewable sources.\n"
- "5. Flexibility: Renewable energy sources are often more flexible and can be adapted to different "
- "situations and needs, while non-renewable sources are more rigid and inflexible.\n"
- "6. Sustainability: Renewable energy sources are more sustainable over the long term, while "
- "non-renewable sources are not, and their depletion can lead to economic and social instability.\n")
- ),
- offset=2,
- sep_style=SeparatorStyle.SINGLE,
- sep="###",
-)
-
-conv_vicuna_v1 = Conversation(
- system="A chat between a curious user and an artificial intelligence assistant. "
- "The assistant gives helpful, detailed, and polite answers to the user's questions.",
- roles=("USER", "ASSISTANT"),
- version="v1",
- messages=(),
- offset=0,
- sep_style=SeparatorStyle.TWO,
- sep=" ",
- sep2="",
-)
-
-conv_llama_2 = Conversation(
- system="""You are a helpful, respectful and honest assistant. Always answer as helpfully as possible, while being safe. Your answers should not include any harmful, unethical, racist, sexist, toxic, dangerous, or illegal content. Please ensure that your responses are socially unbiased and positive in nature.
-
-If a question does not make any sense, or is not factually coherent, explain why instead of answering something not correct. If you don't know the answer to a question, please don't share false information.""",
- roles=("USER", "ASSISTANT"),
- version="llama_v2",
- messages=(),
- offset=0,
- sep_style=SeparatorStyle.LLAMA_2,
- sep="",
- sep2="",
-)
-
-conv_llava_llama_2 = Conversation(
- system="You are a helpful language and vision assistant. "
- "You are able to understand the visual content that the user provides, "
- "and assist the user with a variety of tasks using natural language.",
- roles=("USER", "ASSISTANT"),
- version="llama_v2",
- messages=(),
- offset=0,
- sep_style=SeparatorStyle.LLAMA_2,
- sep="",
- sep2="",
-)
-
-conv_mpt = Conversation(
- system="""<|im_start|>system
-A conversation between a user and an LLM-based AI assistant. The assistant gives helpful and honest answers.""",
- roles=("<|im_start|>user\n", "<|im_start|>assistant\n"),
- version="mpt",
- messages=(),
- offset=0,
- sep_style=SeparatorStyle.MPT,
- sep="<|im_end|>",
-)
-
-conv_llava_plain = Conversation(
- system="",
- roles=("", ""),
- messages=(
- ),
- offset=0,
- sep_style=SeparatorStyle.PLAIN,
- sep="\n",
-)
-
-conv_llava_v0 = Conversation(
- system="A chat between a curious human and an artificial intelligence assistant. "
- "The assistant gives helpful, detailed, and polite answers to the human's questions.",
- roles=("Human", "Assistant"),
- messages=(
- ),
- offset=0,
- sep_style=SeparatorStyle.SINGLE,
- sep="###",
-)
-
-conv_llava_v0_mmtag = Conversation(
- system="A chat between a curious user and an artificial intelligence assistant. "
- "The assistant is able to understand the visual content that the user provides, and assist the user with a variety of tasks using natural language."
- "The visual content will be provided with the following format: visual content.",
- roles=("Human", "Assistant"),
- messages=(
- ),
- offset=0,
- sep_style=SeparatorStyle.SINGLE,
- sep="###",
- version="v0_mmtag",
-)
-
-conv_llava_v1 = Conversation(
- system="A chat between a curious human and an artificial intelligence assistant. "
- "The assistant gives helpful, detailed, and polite answers to the human's questions.",
- roles=("USER", "ASSISTANT"),
- version="v1",
- messages=(),
- offset=0,
- sep_style=SeparatorStyle.TWO,
- sep=" ",
- sep2="",
-)
-
-conv_llava_v1_mmtag = Conversation(
- system="A chat between a curious user and an artificial intelligence assistant. "
- "The assistant is able to understand the visual content that the user provides, and assist the user with a variety of tasks using natural language."
- "The visual content will be provided with the following format: visual content.",
- roles=("USER", "ASSISTANT"),
- messages=(),
- offset=0,
- sep_style=SeparatorStyle.TWO,
- sep=" ",
- sep2="",
- version="v1_mmtag",
-)
-
-conv_mistral_instruct = Conversation(
- system="",
- roles=("USER", "ASSISTANT"),
- version="llama_v2",
- messages=(),
- offset=0,
- sep_style=SeparatorStyle.LLAMA_2,
- sep="",
- sep2="",
-)
-
-default_conversation = conv_vicuna_v1
-conv_templates = {
- "default": conv_vicuna_v0,
- "v0": conv_vicuna_v0,
- "v1": conv_vicuna_v1,
- "vicuna_v1": conv_vicuna_v1,
- "llama_2": conv_llama_2,
- "mistral_instruct": conv_mistral_instruct,
-
- "plain": conv_llava_plain,
- "v0_plain": conv_llava_plain,
- "llava_v0": conv_llava_v0,
- "v0_mmtag": conv_llava_v0_mmtag,
- "llava_v1": conv_llava_v1,
- "v1_mmtag": conv_llava_v1_mmtag,
- "llava_llama_2": conv_llava_llama_2,
- "mpt": conv_mpt,
-}
-
-
-if __name__ == "__main__":
- print(default_conversation.get_prompt())
diff --git a/LLaVA-Med/llava/eval/eval_multimodal_chat_gpt_score.py b/LLaVA-Med/llava/eval/eval_multimodal_chat_gpt_score.py
deleted file mode 100644
index 31bdf95ddab53987b24c69f58faf25b0ea6314cf..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/eval/eval_multimodal_chat_gpt_score.py
+++ /dev/null
@@ -1,112 +0,0 @@
-import os
-import json
-import argparse
-from copy import deepcopy
-import itertools
-from typing import Any
-from operator import add
-from pprint import pprint
-from typing import List
-from pathlib import Path
-from tqdm import tqdm
-
-import llm
-import util
-
-
-INSTRUCT_PROMPT = """We would like to request your feedback on the performance of two AI assistants in response to the user question displayed above. The user asks the question on observing an image. For your reference, the visual content in the image is represented with caption describing the same image.
- Please rate the helpfulness, relevance, accuracy, level of details of their responses. Each assistant receives an overall score on a scale of 1 to 10, where a higher score indicates better overall performance.
- Please first output a single line containing only two values indicating the scores for Assistant 1 and 2, respectively. The two scores are separated by a space. In the subsequent line, please provide a comprehensive explanation of your evaluation, avoiding any potential bias and ensuring that the order in which the responses were presented does not affect your judgment."""
-ROLE = 'Assistant'
-
-# Generate instruction for GPT-4 to score the two answers.
-def conv_to_str(fig_label, fig_caption, fig_context, question, ans1, ans2):
- return (f'[Context]\n'
- f'Figure Caption:\n{fig_label}: {fig_caption}\n\n'
- f'Figure Context:\n\t- {fig_context}\n\n'
- f'[Question]\n{question}\n\n'
- f'[{ROLE} 1]\n{ans1}\n\n[End of {ROLE} 1]\n\n'
- f'[{ROLE} 2]\n{ans2}\n\n[End of {ROLE} 2]\n\n'
- f'[System]\n{INSTRUCT_PROMPT}\n\n')
-
-def compare_messages_gen(fig_label, fig_caption, fig_context, question, ans1, ans2):
- messages = [
- {"role": "system", "content": """'You are a helpful and precise assistant for checking the quality of the answer."""},
- ]
- messages.append({"role": "user", "content": conv_to_str(fig_label, fig_caption, fig_context, question, ans1, ans2)})
- return messages
-
-
-def sum_list_list(x):
- return sum(item for inner_list in x for item in inner_list)
-
-def chunk(lst, n):
- for i in range(0, len(lst), n):
- if i+(1.5*n)=BATCH_SIZE:
- inference_results = [x.strip() for chunk_messages in chunk([x for x in batch if x], BATCH_SIZE) for x in model_inst.infer(chunk_messages)]
- for item, inference_result in zip(batch_samples, inference_results):
- item['gpt_eval'] = inference_result
- results.extend(batch_samples)
- batch = []
- batch_samples = []
- inference_results = [x.strip() for chunk_messages in chunk([x for x in batch if x], BATCH_SIZE) for x in model_inst.infer(chunk_messages)]
- for item, inference_result in zip(batch_samples, inference_results):
- item['gpt_eval'] = inference_result
- results.extend(batch_samples)
- print(f"Result Size: {len(results)}")
- return results
-
-
-def main(args):
- answer_data = util.load_file_jsonl(args.answers_file)
- question_data = util.load_file_jsonl(args.question_file)
-
- samples = []
- for question, answer in zip(question_data, answer_data):
- question_copy = deepcopy(question)
- question['question'] = question_copy['text']
- question['ans1'] = question_copy.pop('gpt4_answer')
- question['ans2'] = answer['text']
- samples.append(question)
-
- results = infer(samples)
-
- # Create parent directory of output score files if it doesn't exist
- os.makedirs(Path(args.scores_file).parent, exist_ok=True)
-
- with open(args.scores_file, 'w') as f:
- for row in results:
- f.write(json.dumps(row)+'\n')
-
-
-if __name__ == '__main__':
- parser = argparse.ArgumentParser("GPT-4 Multimodal Chat Scoring", add_help=True)
- parser.add_argument("--answers-file", default="", metavar="FILE", help="path to model answer file")
- parser.add_argument("--question-file", default="data/questions/llava_med_eval_qa50_qa.jsonl", metavar="FILE", help="path to multichat questions file")
- parser.add_argument("--scores-file", default="", metavar="FILE", help="path to save gpt-4 score file")
- args = parser.parse_args()
- main(args)
\ No newline at end of file
diff --git a/LLaVA-Med/llava/eval/llm.py b/LLaVA-Med/llava/eval/llm.py
deleted file mode 100644
index 2394f3cd459f575479fffd1fb7ff1a08068bb332..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/eval/llm.py
+++ /dev/null
@@ -1,134 +0,0 @@
-import os
-import abc
-import asyncio
-from abc import abstractmethod
-import math
-
-import tiktoken
-import openai
-import backoff
-
-
-class LLM(abc.ABC):
-
- prompt_percent = 0.9
-
- @abstractmethod
- def __init__(self):
- raise NotImplementedError("Subclasses should implement this!")
-
- @abstractmethod
- def infer(self, prompts):
- raise NotImplementedError("Subclasses should implement this!")
-
- @abstractmethod
- def split_input(self, fixed_instruction, few_shot_examples, splittable_input, input_header, output_header):
- raise NotImplementedError("Subclasses should implement this!")
-
-
-class GPT(LLM):
-
- prompt_percent = 0.8
-
- openai_cxn_dict = {
- 'default': {
- 'endpoint': "INSERT YOUR AZURE OPENAI ENDPOINT HERE",
- 'api_key': "INSERT YOUR AZURE OPENAI API KEY HERE",
- },
- }
-
- deployment_max_length_dict = {
- 'gpt-4': 8192,
- 'gpt-4-0314': 8192,
- 'gpt-4-32k': 32768,
- 'gpt-35-turbo': 4096,
- 'gpt-35-turbo-16k': 16385,
- }
-
- def __init__(self, model_id):
- self.temperature = 0.0
- self.top_k = 1
- self.encoding = tiktoken.encoding_for_model("-".join(model_id.split("-", 2)[:2]).replace('5', '.5'))
- self.openai_api = 'default'
- self.model_id = model_id
- self.max_length = self.deployment_max_length_dict[model_id]
- self.client = openai.AsyncAzureOpenAI(
- api_key=self.openai_cxn_dict[self.openai_api]['api_key'],
- api_version="2023-12-01-preview",
- azure_endpoint=self.openai_cxn_dict[self.openai_api]['endpoint']
- )
-
- def gen_messages(self, fixed_instruction, few_shot_examples, input, input_header, output_header):
- messages = [
- {
- "role": "system",
- "content": fixed_instruction,
- },
- ]
- for example in few_shot_examples:
- messages.extend(
- [
- {
- "role": "user",
- "content": input_header+'\n'+example['user']+'\n\n'+output_header,
- },
- {
- "role": "assistant",
- "content": example['assistant'],
- },
- ]
- )
- messages.extend(
- [
- {
- "role": "user",
- "content": input_header+'\n'+input+'\n\n'+output_header,
- },
- ]
- )
- return messages
-
- # Define the coroutine for making API calls to GPT
- @backoff.on_exception(backoff.expo, openai.RateLimitError)
- async def make_api_call_to_gpt(
- self,
- messages
- ):
- response = await self.client.chat.completions.create(
- model=self.model_id,
- messages=messages,
- temperature=self.temperature,
- )
- return response.choices[0].message.content
-
- async def dispatch_openai_requests(
- self,
- messages_list,
- ):
- # Asynchronously call the function for each prompt
- tasks = [self.make_api_call_to_gpt(messages) for messages in messages_list]
-
- # Gather and run the tasks concurrently
- results = await asyncio.gather(*tasks)
- return results
-
- def infer(self,
- messages_list,
- ):
- return asyncio.run(self.dispatch_openai_requests(messages_list))
-
- def split_input(self, fixed_instruction, few_shot_examples, splittable_input, input_header, output_header):
- # Tokenize fixed_prompt
- fixed_token_ids = self.encoding.encode(fixed_instruction+' '.join([x['user']+' '+x['assistant'] for x in few_shot_examples]))
- # Calculate remaining token length
- remaining_token_len = math.ceil((self.prompt_percent*self.max_length)-len(fixed_token_ids))
-
- # Tokenize splittable_input
- split_token_ids = self.encoding.encode(splittable_input)
-
- # Split tokenized split_prompt into list of individual inputs strings. Uses tokens to calculate length
- split_token_ids_list = [split_token_ids[i:i+remaining_token_len+10] for i in range(0, len(split_token_ids), remaining_token_len)]
- split_input_list = [self.encoding.decode(split_token_ids) for split_token_ids in split_token_ids_list]
-
- # Take the fixed_prompt, few_shot_examples, splitted inputs, and input/output headers and generate list of prompt strings.
- return [self.gen_messages(fixed_instruction, few_shot_examples, split_input, input_header, output_header) for split_input in split_input_list]
\ No newline at end of file
diff --git a/LLaVA-Med/llava/eval/model_vqa.py b/LLaVA-Med/llava/eval/model_vqa.py
deleted file mode 100644
index 4f7a09eb1f5dffded5c21ed756136c3a69c9603e..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/eval/model_vqa.py
+++ /dev/null
@@ -1,109 +0,0 @@
-import argparse
-import torch
-import os
-import json
-from tqdm import tqdm
-import shortuuid
-
-from llava.constants import IMAGE_TOKEN_INDEX, DEFAULT_IMAGE_TOKEN, DEFAULT_IM_START_TOKEN, DEFAULT_IM_END_TOKEN
-from llava.conversation import conv_templates, SeparatorStyle
-from llava.model.builder import load_pretrained_model
-from llava.utils import disable_torch_init
-from llava.mm_utils import tokenizer_image_token, get_model_name_from_path, KeywordsStoppingCriteria, process_images
-
-from PIL import Image
-import math
-from transformers import set_seed, logging
-
-logging.set_verbosity_error()
-
-
-def split_list(lst, n):
- """Split a list into n (roughly) equal-sized chunks"""
- chunk_size = math.ceil(len(lst) / n) # integer division
- return [lst[i:i+chunk_size] for i in range(0, len(lst), chunk_size)]
-
-
-def get_chunk(lst, n, k):
- chunks = split_list(lst, n)
- return chunks[k]
-
-
-def eval_model(args):
- set_seed(0)
- # Model
- disable_torch_init()
- model_path = os.path.expanduser(args.model_path)
- model_name = get_model_name_from_path(model_path)
- tokenizer, model, image_processor, context_len = load_pretrained_model(model_path, args.model_base, model_name)
-
- questions = [json.loads(q) for q in open(os.path.expanduser(args.question_file), "r")]
- questions = get_chunk(questions, args.num_chunks, args.chunk_idx)
- answers_file = os.path.expanduser(args.answers_file)
- os.makedirs(os.path.dirname(answers_file), exist_ok=True)
- ans_file = open(answers_file, "w")
- for line in tqdm(questions):
- idx = line["question_id"]
- image_file = line["image"]
- qs = line["text"].replace(DEFAULT_IMAGE_TOKEN, '').strip()
- cur_prompt = qs
- if model.config.mm_use_im_start_end:
- qs = DEFAULT_IM_START_TOKEN + DEFAULT_IMAGE_TOKEN + DEFAULT_IM_END_TOKEN + '\n' + qs
- else:
- qs = DEFAULT_IMAGE_TOKEN + '\n' + qs
-
- conv = conv_templates[args.conv_mode].copy()
- conv.append_message(conv.roles[0], qs)
- conv.append_message(conv.roles[1], None)
- prompt = conv.get_prompt()
-
- input_ids = tokenizer_image_token(prompt, tokenizer, IMAGE_TOKEN_INDEX, return_tensors='pt').unsqueeze(0).cuda()
-
- image = Image.open(os.path.join(args.image_folder, image_file))
- image_tensor = process_images([image], image_processor, model.config)[0]
-
- stop_str = conv.sep if conv.sep_style != SeparatorStyle.TWO else conv.sep2
- keywords = [stop_str]
- stopping_criteria = KeywordsStoppingCriteria(keywords, tokenizer, input_ids)
-
- with torch.inference_mode():
- output_ids = model.generate(
- input_ids,
- images=image_tensor.unsqueeze(0).half().cuda(),
- do_sample=True if args.temperature > 0 else False,
- temperature=args.temperature,
- top_p=args.top_p,
- num_beams=args.num_beams,
- # no_repeat_ngram_size=3,
- max_new_tokens=1024,
- use_cache=True)
-
- outputs = tokenizer.batch_decode(output_ids, skip_special_tokens=True)[0].strip()
-
- ans_id = shortuuid.uuid()
- ans_file.write(json.dumps({"question_id": idx,
- "prompt": cur_prompt,
- "text": outputs,
- "answer_id": ans_id,
- "model_id": model_name,
- "metadata": {}}) + "\n")
- ans_file.flush()
- ans_file.close()
-
-
-if __name__ == "__main__":
- parser = argparse.ArgumentParser()
- parser.add_argument("--model-path", type=str, default="facebook/opt-350m")
- parser.add_argument("--model-base", type=str, default=None)
- parser.add_argument("--image-folder", type=str, default="")
- parser.add_argument("--question-file", type=str, default="tables/question.jsonl")
- parser.add_argument("--answers-file", type=str, default="answer.jsonl")
- parser.add_argument("--conv-mode", type=str, default="vicuna_v1")
- parser.add_argument("--num-chunks", type=int, default=1)
- parser.add_argument("--chunk-idx", type=int, default=0)
- parser.add_argument("--temperature", type=float, default=0.2)
- parser.add_argument("--top_p", type=float, default=None)
- parser.add_argument("--num_beams", type=int, default=1)
- args = parser.parse_args()
-
- eval_model(args)
diff --git a/LLaVA-Med/llava/eval/run_llava.py b/LLaVA-Med/llava/eval/run_llava.py
deleted file mode 100644
index 24b0fffcc11a2045dfc7f5ac6cae4f057aaba6d6..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/eval/run_llava.py
+++ /dev/null
@@ -1,145 +0,0 @@
-import argparse
-import torch
-
-from llava.constants import (
- IMAGE_TOKEN_INDEX,
- DEFAULT_IMAGE_TOKEN,
- DEFAULT_IM_START_TOKEN,
- DEFAULT_IM_END_TOKEN,
- IMAGE_PLACEHOLDER,
-)
-from llava.conversation import conv_templates, SeparatorStyle
-from llava.model.builder import load_pretrained_model
-from llava.utils import disable_torch_init
-from llava.mm_utils import (
- process_images,
- tokenizer_image_token,
- get_model_name_from_path,
-)
-
-from PIL import Image
-
-import requests
-from PIL import Image
-from io import BytesIO
-import re
-
-
-def image_parser(args):
- out = args.image_file.split(args.sep)
- return out
-
-
-def load_image(image_file):
- if image_file.startswith("http") or image_file.startswith("https"):
- response = requests.get(image_file)
- image = Image.open(BytesIO(response.content)).convert("RGB")
- else:
- image = Image.open(image_file).convert("RGB")
- return image
-
-
-def load_images(image_files):
- out = []
- for image_file in image_files:
- image = load_image(image_file)
- out.append(image)
- return out
-
-
-def eval_model(args):
- # Model
- disable_torch_init()
-
- model_name = get_model_name_from_path(args.model_path)
- tokenizer, model, image_processor, context_len = load_pretrained_model(
- args.model_path, args.model_base, model_name
- )
-
- qs = args.query
- image_token_se = DEFAULT_IM_START_TOKEN + DEFAULT_IMAGE_TOKEN + DEFAULT_IM_END_TOKEN
- if IMAGE_PLACEHOLDER in qs:
- if model.config.mm_use_im_start_end:
- qs = re.sub(IMAGE_PLACEHOLDER, image_token_se, qs)
- else:
- qs = re.sub(IMAGE_PLACEHOLDER, DEFAULT_IMAGE_TOKEN, qs)
- else:
- if model.config.mm_use_im_start_end:
- qs = image_token_se + "\n" + qs
- else:
- qs = DEFAULT_IMAGE_TOKEN + "\n" + qs
-
- if "llama-2" in model_name.lower():
- conv_mode = "llava_llama_2"
- elif "mistral" in model_name.lower():
- conv_mode = "mistral_instruct"
- elif "v1.6-34b" in model_name.lower():
- conv_mode = "chatml_direct"
- elif "v1" in model_name.lower():
- conv_mode = "llava_v1"
- elif "mpt" in model_name.lower():
- conv_mode = "mpt"
- else:
- conv_mode = "llava_v0"
-
- if args.conv_mode is not None and conv_mode != args.conv_mode:
- print(
- "[WARNING] the auto inferred conversation mode is {}, while `--conv-mode` is {}, using {}".format(
- conv_mode, args.conv_mode, args.conv_mode
- )
- )
- else:
- args.conv_mode = conv_mode
-
- conv = conv_templates[args.conv_mode].copy()
- conv.append_message(conv.roles[0], qs)
- conv.append_message(conv.roles[1], None)
- prompt = conv.get_prompt()
-
- image_files = image_parser(args)
- images = load_images(image_files)
- image_sizes = [x.size for x in images]
- images_tensor = process_images(
- images,
- image_processor,
- model.config
- ).to(model.device, dtype=torch.float16)
-
- input_ids = (
- tokenizer_image_token(prompt, tokenizer, IMAGE_TOKEN_INDEX, return_tensors="pt")
- .unsqueeze(0)
- .cuda()
- )
-
- with torch.inference_mode():
- output_ids = model.generate(
- input_ids,
- images=images_tensor,
- image_sizes=image_sizes,
- do_sample=True if args.temperature > 0 else False,
- temperature=args.temperature,
- top_p=args.top_p,
- num_beams=args.num_beams,
- max_new_tokens=args.max_new_tokens,
- use_cache=True,
- )
-
- outputs = tokenizer.batch_decode(output_ids, skip_special_tokens=True)[0].strip()
- print(outputs)
-
-
-if __name__ == "__main__":
- parser = argparse.ArgumentParser()
- parser.add_argument("--model-path", type=str, default="facebook/opt-350m")
- parser.add_argument("--model-base", type=str, default=None)
- parser.add_argument("--image-file", type=str, required=True)
- parser.add_argument("--query", type=str, required=True)
- parser.add_argument("--conv-mode", type=str, default=None)
- parser.add_argument("--sep", type=str, default=",")
- parser.add_argument("--temperature", type=float, default=0.2)
- parser.add_argument("--top_p", type=float, default=None)
- parser.add_argument("--num_beams", type=int, default=1)
- parser.add_argument("--max_new_tokens", type=int, default=512)
- args = parser.parse_args()
-
- eval_model(args)
diff --git a/LLaVA-Med/llava/eval/summarize_gpt_review.py b/LLaVA-Med/llava/eval/summarize_gpt_review.py
deleted file mode 100644
index 971961b4e8e9a7edf329b54a63e0b0b2096fa2e9..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/eval/summarize_gpt_review.py
+++ /dev/null
@@ -1,47 +0,0 @@
-import argparse
-from copy import deepcopy
-import util
-from pprint import pprint
-from collections import defaultdict
-import pandas as pd
-import json
-
-
-def get_domain(x):
- for domain in ['chest_xray', 'mri', 'histology', 'gross', 'ct_scan']:
- in_domain = x['domain'][domain]
- if in_domain:
- return domain
-
-
-
-def main(args):
- scores_data = util.load_file_jsonl(args.scores_file)
- predictions = [(x['question_id'], x['type'], get_domain(x), x['gpt_eval'].split('\n')[0].split(' ')) for x in scores_data]
-
- score_type_dict = defaultdict(lambda: defaultdict(list))
- for q_id, q_type, domain, (a1_score, a2_score) in predictions:
- score_type_dict[q_type][1].append(a1_score)
- score_type_dict[q_type][2].append(a2_score)
- score_type_dict['overall'][1].append(a1_score)
- score_type_dict['overall'][2].append(a2_score)
- score_type_dict[domain][1].append(a1_score)
- score_type_dict[domain][2].append(a2_score)
-
- result = defaultdict(dict)
-
- for q_type, score_dict in score_type_dict.items():
- result[q_type]['gpt4_score'] = util.get_avg(score_dict[1])
- result[q_type]['pred_score'] = util.get_avg(score_dict[2])
- result[q_type]['pred_relative_score'] = util.get_avg([float(s2)/float(s1) for s1, s2 in zip(score_dict[1], score_dict[2])])*100
- result[q_type]['data_size'] = len(score_dict[1])
-
- df = pd.DataFrame.from_dict(result).filter(['conversation', 'detailed_description', 'chest_xray', 'mri', 'histology', 'gross', 'ct_scan', 'overall'])
- print(df)
-
-
-if __name__ == '__main__':
- parser = argparse.ArgumentParser("GPT-4 Multimodal Chat Eval Postprocessing", add_help=True)
- parser.add_argument("--scores-file", default="", metavar="FILE", help="input path to gpt-4 score file")
- args = parser.parse_args()
- main(args)
\ No newline at end of file
diff --git a/LLaVA-Med/llava/eval/util.py b/LLaVA-Med/llava/eval/util.py
deleted file mode 100644
index a245850bb5f00f696b741491637d3e92f23d471c..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/eval/util.py
+++ /dev/null
@@ -1,9 +0,0 @@
-import json
-
-
-def load_file_jsonl(path):
- with open(path) as f:
- return [json.loads(row) for row in f]
-
-def get_avg(x):
- return sum([float(y) for y in x])/len(x)
\ No newline at end of file
diff --git a/LLaVA-Med/llava/mm_utils.py b/LLaVA-Med/llava/mm_utils.py
deleted file mode 100644
index 2a6f93fcf1941e95001b59f20b447cd03667173a..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/mm_utils.py
+++ /dev/null
@@ -1,110 +0,0 @@
-from PIL import Image
-from io import BytesIO
-import base64
-import random
-import torch
-from transformers import StoppingCriteria
-from llava.constants import IMAGE_TOKEN_INDEX
-
-
-def load_image_from_base64(image):
- return Image.open(BytesIO(base64.b64decode(image)))
-
-
-def expand2square(pil_img, background_color):
- width, height = pil_img.size
- if width == height:
- return pil_img
- elif width > height:
- result = Image.new(pil_img.mode, (width, width), background_color)
- # sample a random between 0 and (width - height) // 2
- y_start = random.randint((width - height) // 2, (width - height) // 2 + 1)
- result.paste(pil_img, (0, y_start))
- return result
- else:
- result = Image.new(pil_img.mode, (height, height), background_color)
- # sample a random between 0 and (height - width) // 2
- x_start = random.randint((height - width) // 2, (height - width) // 2 + 1)
- result.paste(pil_img, (x_start, 0))
- return result
-
-
-def process_images(images, image_processor, model_cfg):
- image_aspect_ratio = getattr(model_cfg, "image_aspect_ratio", None)
- new_images = []
- for image in images:
- if image_aspect_ratio == 'pad':
- if image.mode=='L':
- background_color = int(255*sum(image_processor.image_mean)/len(image_processor.image_mean))
- else:
- background_color = tuple(int(x*255) for x in image_processor.image_mean)
- image = expand2square(image, background_color)
- image = image_processor.preprocess(image, return_tensors='pt')['pixel_values'][0]
- new_images.append(image)
- if all(x.shape == new_images[0].shape for x in new_images):
- new_images = torch.stack(new_images, dim=0)
- return new_images
-
-
-def tokenizer_image_token(prompt, tokenizer, image_token_index=IMAGE_TOKEN_INDEX, return_tensors=None):
- prompt_chunks = [tokenizer(chunk).input_ids for chunk in prompt.split('')]
-
- def insert_separator(X, sep):
- return [ele for sublist in zip(X, [sep]*len(X)) for ele in sublist][:-1]
-
- input_ids = []
- offset = 0
- if len(prompt_chunks) > 0 and len(prompt_chunks[0]) > 0 and prompt_chunks[0][0] == tokenizer.bos_token_id:
- offset = 1
- input_ids.append(prompt_chunks[0][0])
-
- for x in insert_separator(prompt_chunks, [image_token_index] * (offset + 1)):
- input_ids.extend(x[offset:])
-
- if return_tensors is not None:
- if return_tensors == 'pt':
- return torch.tensor(input_ids, dtype=torch.long)
- raise ValueError(f'Unsupported tensor type: {return_tensors}')
- return input_ids
-
-
-def get_model_name_from_path(model_path):
- model_path = model_path.strip("/")
- model_paths = model_path.split("/")
- if model_paths[-1].startswith('checkpoint-'):
- return model_paths[-2] + "_" + model_paths[-1]
- else:
- return model_paths[-1]
-
-class KeywordsStoppingCriteria(StoppingCriteria):
- def __init__(self, keywords, tokenizer, input_ids):
- self.keywords = keywords
- self.keyword_ids = []
- self.max_keyword_len = 0
- for keyword in keywords:
- cur_keyword_ids = tokenizer(keyword).input_ids
- if len(cur_keyword_ids) > 1 and cur_keyword_ids[0] == tokenizer.bos_token_id:
- cur_keyword_ids = cur_keyword_ids[1:]
- if len(cur_keyword_ids) > self.max_keyword_len:
- self.max_keyword_len = len(cur_keyword_ids)
- self.keyword_ids.append(torch.tensor(cur_keyword_ids))
- self.tokenizer = tokenizer
- self.start_len = input_ids.shape[1]
-
- def call_for_batch(self, output_ids: torch.LongTensor, scores: torch.FloatTensor, **kwargs) -> bool:
- offset = min(output_ids.shape[1] - self.start_len, self.max_keyword_len)
- self.keyword_ids = [keyword_id.to(output_ids.device) for keyword_id in self.keyword_ids]
- for keyword_id in self.keyword_ids:
- if (output_ids[0, -keyword_id.shape[0]:] == keyword_id).all():
- return True
- outputs = self.tokenizer.batch_decode(output_ids[:, -offset:], skip_special_tokens=True)[0]
- for keyword in self.keywords:
- if keyword in outputs:
- return True
- return False
-
- def __call__(self, output_ids: torch.LongTensor, scores: torch.FloatTensor, **kwargs) -> bool:
- outputs = []
- for i in range(output_ids.shape[0]):
- outputs.append(self.call_for_batch(output_ids[i].unsqueeze(0), scores))
- return all(outputs)
diff --git a/LLaVA-Med/llava/model/__init__.py b/LLaVA-Med/llava/model/__init__.py
deleted file mode 100644
index 896bc6b698efbc3c5dc20d06f29e7d57d6a43672..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/model/__init__.py
+++ /dev/null
@@ -1 +0,0 @@
-from .language_model.llava_mistral import LlavaMistralForCausalLM, LlavaMistralConfig
diff --git a/LLaVA-Med/llava/model/builder.py b/LLaVA-Med/llava/model/builder.py
deleted file mode 100644
index 187020cfcadbee358e6e577b4289a477c9011ddb..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/model/builder.py
+++ /dev/null
@@ -1,83 +0,0 @@
-from transformers import AutoTokenizer, AutoModelForCausalLM, AutoConfig, BitsAndBytesConfig
-import torch
-from llava.model import LlavaMistralForCausalLM
-from llava.constants import DEFAULT_IMAGE_PATCH_TOKEN, DEFAULT_IM_START_TOKEN, DEFAULT_IM_END_TOKEN
-
-
-def load_pretrained_model(model_path, model_base, model_name, load_8bit=False, load_4bit=False, device_map="auto", device="cuda"):
-
- kwargs = {}
-
- if device != "cuda":
- kwargs['device_map'] = {"": device}
-
- if load_8bit:
- kwargs['load_in_8bit'] = True
- elif load_4bit:
- kwargs['load_in_4bit'] = True
- kwargs['quantization_config'] = BitsAndBytesConfig(
- load_in_4bit=True,
- bnb_4bit_compute_dtype=torch.float16,
- bnb_4bit_use_double_quant=True,
- bnb_4bit_quant_type='nf4'
- )
- else:
- kwargs['torch_dtype'] = torch.float16
-
- if 'llava' in model_name.lower():
- # Load LLaVA model
- if 'mistral' in model_name.lower():
- tokenizer = AutoTokenizer.from_pretrained(model_path)
- model = LlavaMistralForCausalLM.from_pretrained(
- model_path,
- low_cpu_mem_usage=False,
- use_flash_attention_2=False,
- **kwargs
- )
- else:
- # Load language model
- if model_base is not None:
- # PEFT model
- from peft import PeftModel
- tokenizer = AutoTokenizer.from_pretrained(model_base, use_fast=False)
- model = AutoModelForCausalLM.from_pretrained(model_base, low_cpu_mem_usage=True, **kwargs)
- print(f"Loading LoRA weights from {model_path}")
- model = PeftModel.from_pretrained(model, model_path)
- print(f"Merging weights")
- model = model.merge_and_unload()
- print('Convert to FP16...')
- model.to(torch.float16)
- else:
- use_fast = False
- if 'mpt' in model_name.lower():
- tokenizer = AutoTokenizer.from_pretrained(model_path, use_fast=True)
- model = AutoModelForCausalLM.from_pretrained(model_path, low_cpu_mem_usage=True, trust_remote_code=True, **kwargs)
- else:
- tokenizer = AutoTokenizer.from_pretrained(model_path, use_fast=False)
- model = AutoModelForCausalLM.from_pretrained(model_path, low_cpu_mem_usage=True, **kwargs)
-
- image_processor = None
-
- if 'llava' in model_name.lower(): # or 'mistral' in model_name.lower():
- mm_use_im_start_end = getattr(model.config, "mm_use_im_start_end", False)
- mm_use_im_patch_token = getattr(model.config, "mm_use_im_patch_token", True)
- if mm_use_im_patch_token:
- tokenizer.add_tokens([DEFAULT_IMAGE_PATCH_TOKEN], special_tokens=True)
- if mm_use_im_start_end:
- tokenizer.add_tokens([DEFAULT_IM_START_TOKEN, DEFAULT_IM_END_TOKEN], special_tokens=True)
- model.resize_token_embeddings(len(tokenizer))
-
- vision_tower = model.get_vision_tower()
- if not vision_tower.is_loaded:
- vision_tower.load_model()
- vision_tower.to(device=device, dtype=torch.float16)
- model.model.mm_projector.to(device=device, dtype=torch.float16)
- model.to(device=device, dtype=torch.float16)
- image_processor = vision_tower.image_processor
-
- if hasattr(model.config, "max_sequence_length"):
- context_len = model.config.max_sequence_length
- else:
- context_len = 2048
-
- return tokenizer, model, image_processor, context_len
diff --git a/LLaVA-Med/llava/model/builders.py b/LLaVA-Med/llava/model/builders.py
deleted file mode 100644
index 923de04d0fcf30181f45dbd3da42b1c284794ff8..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/model/builders.py
+++ /dev/null
@@ -1,152 +0,0 @@
-import os
-import warnings
-import shutil
-
-from transformers import AutoTokenizer, AutoModelForCausalLM, AutoConfig, BitsAndBytesConfig
-import torch
-from llava.model import LLavaMistralForCausalLM
-from llava.constants import DEFAULT_IMAGE_PATCH_TOKEN, DEFAULT_IM_START_TOKEN, DEFAULT_IM_END_TOKEN
-
-
-def load_pretrained_model(model_path, model_base, model_name, load_8bit=False, load_4bit=False, device_map="auto", device="cuda", use_flash_attn=False, **kwargs):
- kwargs = {"device_map": device_map, **kwargs}
-
- if device != "cuda":
- kwargs['device_map'] = {"": device}
-
- if load_8bit:
- kwargs['load_in_8bit'] = True
- elif load_4bit:
- kwargs['load_in_4bit'] = True
- kwargs['quantization_config'] = BitsAndBytesConfig(
- load_in_4bit=True,
- bnb_4bit_compute_dtype=torch.float16,
- bnb_4bit_use_double_quant=True,
- bnb_4bit_quant_type='nf4'
- )
- else:
- kwargs['torch_dtype'] = torch.float16
-
- if use_flash_attn:
- kwargs['attn_implementation'] = 'flash_attention_2'
-
- if 'llava' in model_name.lower():
- # Load LLaVA model
- if 'lora' in model_name.lower() and model_base is None:
- warnings.warn('There is `lora` in model name but no `model_base` is provided. If you are loading a LoRA model, please provide the `model_base` argument. Detailed instruction: https://github.com/haotian-liu/LLaVA#launch-a-model-worker-lora-weights-unmerged.')
- if 'lora' in model_name.lower() and model_base is not None:
- from llava.model.language_model.llava_mistral import LlavaMistralConfig
- lora_cfg_pretrained = LlavaMistralConfig.from_pretrained(model_path)
- tokenizer = AutoTokenizer.from_pretrained(model_base, use_fast=False)
- print('Loading LLaVA from base model...')
- model = LlavaMistralForCausalLM.from_pretrained(model_base, low_cpu_mem_usage=True, config=lora_cfg_pretrained, **kwargs)
- token_num, tokem_dim = model.lm_head.out_features, model.lm_head.in_features
- if model.lm_head.weight.shape[0] != token_num:
- model.lm_head.weight = torch.nn.Parameter(torch.empty(token_num, tokem_dim, device=model.device, dtype=model.dtype))
- model.model.embed_tokens.weight = torch.nn.Parameter(torch.empty(token_num, tokem_dim, device=model.device, dtype=model.dtype))
-
- # print('Loading additional LLaVA weights...')
- # if os.path.exists(os.path.join(model_path, 'non_lora_trainables.bin')):
- # non_lora_trainables = torch.load(os.path.join(model_path, 'non_lora_trainables.bin'), map_location='cpu')
- # else:
- # # this is probably from HF Hub
- # from huggingface_hub import hf_hub_download
- # def load_from_hf(repo_id, filename, subfolder=None):
- # cache_file = hf_hub_download(
- # repo_id=repo_id,
- # filename=filename,
- # subfolder=subfolder)
- # return torch.load(cache_file, map_location='cpu')
- # non_lora_trainables = load_from_hf(model_path, 'non_lora_trainables.bin')
- # non_lora_trainables = {(k[11:] if k.startswith('base_model.') else k): v for k, v in non_lora_trainables.items()}
- # if any(k.startswith('model.model.') for k in non_lora_trainables):
- # non_lora_trainables = {(k[6:] if k.startswith('model.') else k): v for k, v in non_lora_trainables.items()}
- # model.load_state_dict(non_lora_trainables, strict=False)
-
- from peft import PeftModel
- print('Loading LoRA weights...')
- model = PeftModel.from_pretrained(model, model_path)
- print('Merging LoRA weights...')
- model = model.merge_and_unload()
- print('Model is loaded...')
- elif model_base is not None:
- # this may be mm projector only
- print('Loading LLaVA from base model...')
- if 'mpt' in model_name.lower():
- if not os.path.isfile(os.path.join(model_path, 'configuration_mpt.py')):
- shutil.copyfile(os.path.join(model_base, 'configuration_mpt.py'), os.path.join(model_path, 'configuration_mpt.py'))
- tokenizer = AutoTokenizer.from_pretrained(model_base, use_fast=True)
- cfg_pretrained = AutoConfig.from_pretrained(model_path, trust_remote_code=True)
- model = LlavaMptForCausalLM.from_pretrained(model_base, low_cpu_mem_usage=True, config=cfg_pretrained, **kwargs)
- else:
- tokenizer = AutoTokenizer.from_pretrained(model_base, use_fast=False)
- cfg_pretrained = AutoConfig.from_pretrained(model_path)
- model = LlavaLlamaForCausalLM.from_pretrained(model_base, low_cpu_mem_usage=True, config=cfg_pretrained, **kwargs)
-
- mm_projector_weights = torch.load(os.path.join(model_path, 'mm_projector.bin'), map_location='cpu')
- mm_projector_weights = {k: v.to(torch.float16) for k, v in mm_projector_weights.items()}
- model.load_state_dict(mm_projector_weights, strict=False)
- else:
- if 'mpt' in model_name.lower():
- tokenizer = AutoTokenizer.from_pretrained(model_path, use_fast=True)
- model = LlavaMptForCausalLM.from_pretrained(model_path, low_cpu_mem_usage=True, **kwargs)
- elif 'mistral' in model_name.lower():
- tokenizer = AutoTokenizer.from_pretrained(model_path)
- model = LlavaMistralForCausalLM.from_pretrained(
- model_path,
- low_cpu_mem_usage=True,
- **kwargs
- )
- else:
- tokenizer = AutoTokenizer.from_pretrained(model_path, use_fast=False)
- model = LlavaLlamaForCausalLM.from_pretrained(
- model_path,
- low_cpu_mem_usage=True,
- **kwargs
- )
- else:
- # Load language model
- if model_base is not None:
- # PEFT model
- from peft import PeftModel
- tokenizer = AutoTokenizer.from_pretrained(model_base, use_fast=False)
- model = AutoModelForCausalLM.from_pretrained(model_base, low_cpu_mem_usage=True, **kwargs)
- print(f"Loading LoRA weights from {model_path}")
- model = PeftModel.from_pretrained(model, model_path)
- print(f"Merging weights")
- model = model.merge_and_unload()
- print('Convert to FP16...')
- model.to(torch.float16)
- else:
- use_fast = False
- if 'mpt' in model_name.lower():
- tokenizer = AutoTokenizer.from_pretrained(model_path, use_fast=True)
- model = AutoModelForCausalLM.from_pretrained(model_path, low_cpu_mem_usage=True, trust_remote_code=True, **kwargs)
- else:
- tokenizer = AutoTokenizer.from_pretrained(model_path, use_fast=False)
- model = AutoModelForCausalLM.from_pretrained(model_path, low_cpu_mem_usage=True, **kwargs)
-
- image_processor = None
-
- if 'mistral' in model_name.lower():
- mm_use_im_start_end = getattr(model.config, "mm_use_im_start_end", False)
- mm_use_im_patch_token = getattr(model.config, "mm_use_im_patch_token", True)
- if mm_use_im_patch_token:
- tokenizer.add_tokens([DEFAULT_IMAGE_PATCH_TOKEN], special_tokens=True)
- if mm_use_im_start_end:
- tokenizer.add_tokens([DEFAULT_IM_START_TOKEN, DEFAULT_IM_END_TOKEN], special_tokens=True)
- model.resize_token_embeddings(len(tokenizer))
-
- vision_tower = model.get_vision_tower()
- if not vision_tower.is_loaded:
- vision_tower.load_model(device_map=device_map)
- if device_map != 'auto':
- vision_tower.to(device=device_map, dtype=torch.float16)
- image_processor = vision_tower.image_processor
-
- if hasattr(model.config, "max_sequence_length"):
- context_len = model.config.max_sequence_length
- else:
- context_len = 2048
-
- return tokenizer, model, image_processor, context_len
diff --git a/LLaVA-Med/llava/model/language_model/llava_mistral.py b/LLaVA-Med/llava/model/language_model/llava_mistral.py
deleted file mode 100644
index 12692b944af05d92b39fa017a74b60745305894b..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/model/language_model/llava_mistral.py
+++ /dev/null
@@ -1,143 +0,0 @@
-from typing import List, Optional, Tuple, Union
-
-import torch
-import torch.nn as nn
-
-from transformers import AutoConfig, AutoModelForCausalLM, \
- MistralConfig, MistralModel, MistralForCausalLM
-
-from transformers.modeling_outputs import CausalLMOutputWithPast
-from transformers.generation.utils import GenerateOutput
-
-from ..llava_arch import LlavaMetaModel, LlavaMetaForCausalLM
-
-
-class LlavaMistralConfig(MistralConfig):
- model_type = "llava_mistral"
-
-
-class LlavaMistralModel(LlavaMetaModel, MistralModel):
- config_class = LlavaMistralConfig
-
- def __init__(self, config: MistralConfig):
- super(LlavaMistralModel, self).__init__(config)
-
-
-class LlavaMistralForCausalLM(MistralForCausalLM, LlavaMetaForCausalLM):
- config_class = LlavaMistralConfig
-
- def __init__(self, config):
- super(MistralForCausalLM, self).__init__(config)
- self.model = LlavaMistralModel(config)
-
- self.lm_head = nn.Linear(config.hidden_size, config.vocab_size, bias=False)
-
- # Initialize weights and apply final processing
- self.post_init()
-
- def get_model(self):
- return self.model
-
- def forward(
- self,
- input_ids: torch.LongTensor = None,
- attention_mask: Optional[torch.Tensor] = None,
- position_ids: Optional[torch.LongTensor] = None,
- past_key_values: Optional[List[torch.FloatTensor]] = None,
- inputs_embeds: Optional[torch.FloatTensor] = None,
- labels: Optional[torch.LongTensor] = None,
- use_cache: Optional[bool] = None,
- output_attentions: Optional[bool] = None,
- output_hidden_states: Optional[bool] = None,
- images: Optional[torch.FloatTensor] = None,
- image_sizes: Optional[List[List[int]]] = None,
- return_dict: Optional[bool] = None,
- ) -> Union[Tuple, CausalLMOutputWithPast]:
-
- if inputs_embeds is None:
- (
- input_ids,
- position_ids,
- attention_mask,
- past_key_values,
- inputs_embeds,
- labels
- ) = self.prepare_inputs_labels_for_multimodal(
- input_ids,
- position_ids,
- attention_mask,
- past_key_values,
- labels,
- images,
- image_sizes
- )
-
- return super().forward(
- input_ids=input_ids,
- attention_mask=attention_mask,
- position_ids=position_ids,
- past_key_values=past_key_values,
- inputs_embeds=inputs_embeds,
- labels=labels,
- use_cache=use_cache,
- output_attentions=output_attentions,
- output_hidden_states=output_hidden_states,
- return_dict=return_dict
- )
-
- @torch.no_grad()
- def generate(
- self,
- inputs: Optional[torch.Tensor] = None,
- images: Optional[torch.Tensor] = None,
- image_sizes: Optional[torch.Tensor] = None,
- **kwargs,
- ) -> Union[GenerateOutput, torch.LongTensor]:
- position_ids = kwargs.pop("position_ids", None)
- attention_mask = kwargs.pop("attention_mask", None)
- if "inputs_embeds" in kwargs:
- raise NotImplementedError("`inputs_embeds` is not supported")
-
- if images is not None:
- (
- inputs,
- position_ids,
- attention_mask,
- _,
- inputs_embeds,
- _
- ) = self.prepare_inputs_labels_for_multimodal(
- inputs,
- position_ids,
- attention_mask,
- None,
- None,
- images,
- image_sizes=image_sizes
- )
- else:
- inputs_embeds = self.get_model().embed_tokens(inputs)
-
- return super().generate(
- position_ids=position_ids,
- attention_mask=attention_mask,
- inputs_embeds=inputs_embeds,
- **kwargs
- )
-
- def prepare_inputs_for_generation(self, input_ids, past_key_values=None,
- inputs_embeds=None, **kwargs):
- images = kwargs.pop("images", None)
- image_sizes = kwargs.pop("image_sizes", None)
- inputs = super().prepare_inputs_for_generation(
- input_ids, past_key_values=past_key_values, inputs_embeds=inputs_embeds, **kwargs
- )
- if images is not None:
- inputs['images'] = images
- if image_sizes is not None:
- inputs['image_sizes'] = image_sizes
- return inputs
-
-
-AutoConfig.register("llava_mistral", LlavaMistralConfig)
-AutoModelForCausalLM.register(LlavaMistralConfig, LlavaMistralForCausalLM)
diff --git a/LLaVA-Med/llava/model/llava_arch.py b/LLaVA-Med/llava/model/llava_arch.py
deleted file mode 100644
index 433c2f372b69cd7f83e2d85c1a7926dc3dcba33c..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/model/llava_arch.py
+++ /dev/null
@@ -1,309 +0,0 @@
-# Copyright 2023 Haotian Liu
-#
-# Licensed under the Apache License, Version 2.0 (the "License");
-# you may not use this file except in compliance with the License.
-# You may obtain a copy of the License at
-#
-# http://www.apache.org/licenses/LICENSE-2.0
-#
-# Unless required by applicable law or agreed to in writing, software
-# distributed under the License is distributed on an "AS IS" BASIS,
-# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
-# See the License for the specific language governing permissions and
-# limitations under the License.
-
-
-from abc import ABC, abstractmethod
-import os
-from glob import glob
-
-import torch
-import torch.nn as nn
-
-from .multimodal_encoder.builder import build_vision_tower
-from .multimodal_projector.builder import build_vision_projector
-
-from llava.constants import IGNORE_INDEX, IMAGE_TOKEN_INDEX, DEFAULT_IMAGE_PATCH_TOKEN, DEFAULT_IM_START_TOKEN, DEFAULT_IM_END_TOKEN
-
-
-class LlavaMetaModel:
-
- def __init__(self, config):
- super(LlavaMetaModel, self).__init__(config)
-
- if hasattr(config, "mm_vision_tower"):
- self.vision_tower = build_vision_tower(config, delay_load=True)
- self.mm_projector = build_vision_projector(config)
-
- def get_vision_tower(self):
- vision_tower = getattr(self, 'vision_tower', None)
- if type(vision_tower) is list:
- vision_tower = vision_tower[0]
- return vision_tower
-
- def initialize_vision_modules(self, model_args, fsdp=None, embed_tokens=None):
- vision_tower = model_args.vision_tower
- mm_vision_select_layer = model_args.mm_vision_select_layer
- mm_vision_select_feature = model_args.mm_vision_select_feature
- pretrain_mm_mlp_adapter = model_args.pretrain_mm_mlp_adapter
-
- self.config.mm_vision_tower = vision_tower
-
- if self.get_vision_tower() is None:
- vision_tower = build_vision_tower(model_args)
-
- if fsdp is not None and len(fsdp) > 0:
- self.vision_tower = [vision_tower]
- else:
- self.vision_tower = vision_tower
- else:
- if fsdp is not None and len(fsdp) > 0:
- vision_tower = self.vision_tower[0]
- else:
- vision_tower = self.vision_tower
- vision_tower.load_model()
-
- self.config.use_mm_proj = True
- self.config.mm_projector_type = getattr(model_args, 'mm_projector_type', 'linear')
- self.config.mm_hidden_size = vision_tower.hidden_size
- self.config.mm_vision_select_layer = mm_vision_select_layer
- self.config.mm_vision_select_feature = mm_vision_select_feature
-
- # add additional configs for segtok
- self.config.feature_outs = model_args.feature_outs
- self.config.img_size = model_args.img_size
- self.config.vision_backbone = model_args.vision_backbone
- self.config.segtok_posembed = model_args.segtok_posembed
-
- if getattr(self, 'mm_projector', None) is None:
- self.mm_projector = build_vision_projector(self.config)
- else:
- # In case it is frozen by LoRA
- for p in self.mm_projector.parameters():
- p.requires_grad = True
-
- # Initialize last layer in mm_projector with weight=0 and bias=mean(embed_tokens)
- if embed_tokens is not None:
- embed_tokens_weight = embed_tokens.weight.data
- self.mm_projector[-1].weight.data.zero_()
- self.mm_projector[-1].bias.data.copy_(embed_tokens_weight.mean(dim=0))
-
- if pretrain_mm_mlp_adapter is not None:
- def get_w(weights, keyword):
- return {k.split(keyword + '.')[1]: v for k, v in weights.items() if keyword in k}
-
- mm_projector_weights = torch.load(pretrain_mm_mlp_adapter, map_location='cpu')
- self.mm_projector.load_state_dict(get_w(mm_projector_weights, 'mm_projector'))
-
- # also load additional learnable parameters during feature alignment
- checkpoint_folder = os.path.dirname(pretrain_mm_mlp_adapter)
- ckpts = glob(f"{checkpoint_folder}/checkpoint-*", recursive = False)
- if len(ckpts) > 0:
- vision_module_weights = torch.load(f"{ckpts[-1]}/mm_projector.bin", map_location='cpu')
- model_dict = get_w(vision_module_weights, 'vision_tower')
- print(f"Loading vision module weights from {ckpts[-1]}/mm_projector.bin")
- # print keys in model_dict
- print(f"Loaded keys: {model_dict.keys()}")
- self.vision_tower.load_state_dict(model_dict, strict=False)
-
-class LlavaMetaForCausalLM(ABC):
-
- @abstractmethod
- def get_model(self):
- pass
-
- def get_vision_tower(self):
- return self.get_model().get_vision_tower()
-
- def encode_images(self, images):
- image_features = self.get_model().get_vision_tower()(images)
- image_features = self.get_model().mm_projector(image_features)
- return image_features
-
- def prepare_inputs_labels_for_multimodal(
- self, input_ids, position_ids, attention_mask, past_key_values, labels, images, image_sizes=None
- ):
- vision_tower = self.get_vision_tower()
- if vision_tower is None or images is None or input_ids.shape[1] == 1:
- if past_key_values is not None and vision_tower is not None and images is not None and input_ids.shape[1] == 1:
- target_shape = past_key_values[-1][-1].shape[-2] + 1
- attention_mask = torch.cat((attention_mask, torch.ones(
- (attention_mask.shape[0], target_shape - attention_mask.shape[1]),
- dtype=attention_mask.dtype,
- device=attention_mask.device
- )), dim=1)
- position_ids = torch.sum(attention_mask, dim=1).unsqueeze(-1) - 1
- return input_ids, position_ids, attention_mask, past_key_values, None, labels
-
- if type(images) is list or images.ndim == 5:
- concat_images = torch.cat([image for image in images], dim=0)
- image_features = self.encode_images(concat_images)
- split_sizes = [image.shape[0] for image in images]
- image_features = torch.split(image_features, split_sizes, dim=0)
- image_features = [x.flatten(0, 1).to(self.device) for x in image_features]
- else:
- image_features = self.encode_images(images).to(self.device)
-
- # TODO: image start / end is not implemented here to support pretraining.
- if getattr(self.config, 'tune_mm_mlp_adapter', False) and getattr(self.config, 'mm_use_im_start_end', False):
- raise NotImplementedError
-
- # Let's just add dummy tensors if they do not exist,
- # it is a headache to deal with None all the time.
- # But it is not ideal, and if you have a better idea,
- # please open an issue / submit a PR, thanks.
- _labels = labels
- _position_ids = position_ids
- _attention_mask = attention_mask
-
- if attention_mask is None:
- attention_mask = torch.ones_like(input_ids, dtype=torch.bool)
- else:
- attention_mask = attention_mask.bool()
- if position_ids is None:
- position_ids = torch.arange(0, input_ids.shape[1], dtype=torch.long, device=input_ids.device)
-
- if labels is None:
- labels = torch.full_like(input_ids, IGNORE_INDEX)
-
- input_ids = [cur_input_ids[cur_attention_mask] for cur_input_ids, cur_attention_mask in zip(input_ids, attention_mask)]
- labels = [cur_labels[cur_attention_mask] for cur_labels, cur_attention_mask in zip(labels, attention_mask)]
-
- new_input_embeds = []
- new_labels = []
- cur_image_idx = 0
- for batch_idx, cur_input_ids in enumerate(input_ids):
- num_images = (cur_input_ids == IMAGE_TOKEN_INDEX).sum()
- if num_images == 0:
- cur_image_features = image_features[cur_image_idx]
- cur_input_embeds_1 = self.get_model().embed_tokens(cur_input_ids)
- cur_input_embeds = torch.cat([cur_input_embeds_1, cur_image_features[0:0]], dim=0)
- new_input_embeds.append(cur_input_embeds)
- new_labels.append(labels[batch_idx])
- cur_image_idx += 1
- continue
-
- image_token_indices = [-1] + torch.where(cur_input_ids == IMAGE_TOKEN_INDEX)[0].tolist() + [cur_input_ids.shape[0]]
- cur_input_ids_noim = []
- cur_labels = labels[batch_idx]
- cur_labels_noim = []
- for i in range(len(image_token_indices) - 1):
- cur_input_ids_noim.append(cur_input_ids[image_token_indices[i]+1:image_token_indices[i+1]])
- cur_labels_noim.append(cur_labels[image_token_indices[i]+1:image_token_indices[i+1]])
-
- split_sizes = [x.shape[0] for x in cur_labels_noim]
- cur_input_embeds = self.get_model().embed_tokens(torch.cat(cur_input_ids_noim))
- cur_input_embeds_no_im = torch.split(cur_input_embeds, split_sizes, dim=0)
- cur_new_input_embeds = []
- cur_new_labels = []
-
- for i in range(num_images + 1):
- cur_new_input_embeds.append(cur_input_embeds_no_im[i])
- cur_new_labels.append(cur_labels_noim[i])
- if i < num_images:
- cur_image_features = image_features[cur_image_idx]
- cur_image_idx += 1
- cur_new_input_embeds.append(cur_image_features)
- cur_new_labels.append(torch.full((cur_image_features.shape[0],), IGNORE_INDEX, device=cur_labels.device, dtype=cur_labels.dtype))
-
- cur_new_input_embeds = torch.cat(cur_new_input_embeds)
- cur_new_labels = torch.cat(cur_new_labels)
-
- new_input_embeds.append(cur_new_input_embeds)
- new_labels.append(cur_new_labels)
-
- # Truncate sequences to max length as image embeddings can make the sequence longer
- tokenizer_model_max_length = getattr(self.config, 'tokenizer_model_max_length', None)
- if tokenizer_model_max_length is not None:
- new_input_embeds = [x[:tokenizer_model_max_length] for x in new_input_embeds]
- new_labels = [x[:tokenizer_model_max_length] for x in new_labels]
-
- # Combine them
- max_len = max(x.shape[0] for x in new_input_embeds)
- batch_size = len(new_input_embeds)
-
- new_input_embeds_padded = []
- new_labels_padded = torch.full((batch_size, max_len), IGNORE_INDEX, dtype=new_labels[0].dtype, device=new_labels[0].device)
- attention_mask = torch.zeros((batch_size, max_len), dtype=attention_mask.dtype, device=attention_mask.device)
- position_ids = torch.zeros((batch_size, max_len), dtype=position_ids.dtype, device=position_ids.device)
-
- for i, (cur_new_embed, cur_new_labels) in enumerate(zip(new_input_embeds, new_labels)):
- cur_len = cur_new_embed.shape[0]
- if getattr(self.config, 'tokenizer_padding_side', 'right') == "left":
- new_input_embeds_padded.append(torch.cat((
- torch.zeros((max_len - cur_len, cur_new_embed.shape[1]), dtype=cur_new_embed.dtype, device=cur_new_embed.device),
- cur_new_embed
- ), dim=0))
- if cur_len > 0:
- new_labels_padded[i, -cur_len:] = cur_new_labels
- attention_mask[i, -cur_len:] = True
- position_ids[i, -cur_len:] = torch.arange(0, cur_len, dtype=position_ids.dtype, device=position_ids.device)
- else:
- new_input_embeds_padded.append(torch.cat((
- cur_new_embed,
- torch.zeros((max_len - cur_len, cur_new_embed.shape[1]), dtype=cur_new_embed.dtype, device=cur_new_embed.device)
- ), dim=0))
- if cur_len > 0:
- new_labels_padded[i, :cur_len] = cur_new_labels
- attention_mask[i, :cur_len] = True
- position_ids[i, :cur_len] = torch.arange(0, cur_len, dtype=position_ids.dtype, device=position_ids.device)
-
- new_input_embeds = torch.stack(new_input_embeds_padded, dim=0)
-
- if _labels is None:
- new_labels = None
- else:
- new_labels = new_labels_padded
-
- if _attention_mask is None:
- attention_mask = None
- else:
- attention_mask = attention_mask.to(dtype=_attention_mask.dtype)
-
- if _position_ids is None:
- position_ids = None
- return None, position_ids, attention_mask, past_key_values, new_input_embeds, new_labels
-
- def initialize_vision_tokenizer(self, model_args, tokenizer):
- if model_args.mm_use_im_patch_token:
- tokenizer.add_tokens([DEFAULT_IMAGE_PATCH_TOKEN], special_tokens=True)
- self.resize_token_embeddings(len(tokenizer))
-
- if model_args.mm_use_im_start_end:
- num_new_tokens = tokenizer.add_tokens([DEFAULT_IM_START_TOKEN, DEFAULT_IM_END_TOKEN], special_tokens=True)
- self.resize_token_embeddings(len(tokenizer))
-
- if num_new_tokens > 0:
- input_embeddings = self.get_input_embeddings().weight.data
- output_embeddings = self.get_output_embeddings().weight.data
-
- input_embeddings_avg = input_embeddings[:-num_new_tokens].mean(
- dim=0, keepdim=True)
- output_embeddings_avg = output_embeddings[:-num_new_tokens].mean(
- dim=0, keepdim=True)
-
- input_embeddings[-num_new_tokens:] = input_embeddings_avg
- output_embeddings[-num_new_tokens:] = output_embeddings_avg
-
- if model_args.tune_mm_mlp_adapter:
- for p in self.get_input_embeddings().parameters():
- p.requires_grad = True
- for p in self.get_output_embeddings().parameters():
- p.requires_grad = False
-
- if model_args.pretrain_mm_mlp_adapter:
- mm_projector_weights = torch.load(model_args.pretrain_mm_mlp_adapter, map_location='cpu')
- embed_tokens_weight = mm_projector_weights['model.embed_tokens.weight']
- assert num_new_tokens == 2
- if input_embeddings.shape == embed_tokens_weight.shape:
- input_embeddings[-num_new_tokens:] = embed_tokens_weight[-num_new_tokens:]
- elif embed_tokens_weight.shape[0] == num_new_tokens:
- input_embeddings[-num_new_tokens:] = embed_tokens_weight
- else:
- raise ValueError(f"Unexpected embed_tokens_weight shape. Pretrained: {embed_tokens_weight.shape}. Current: {input_embeddings.shape}. Numer of new tokens: {num_new_tokens}.")
- elif model_args.mm_use_im_patch_token:
- if model_args.tune_mm_mlp_adapter:
- for p in self.get_input_embeddings().parameters():
- p.requires_grad = False
- for p in self.get_output_embeddings().parameters():
- p.requires_grad = False
diff --git a/LLaVA-Med/llava/model/multimodal_encoder/builder.py b/LLaVA-Med/llava/model/multimodal_encoder/builder.py
deleted file mode 100644
index cdf0b59b1334ec31e103703ab1693a2accf7b9ac..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/model/multimodal_encoder/builder.py
+++ /dev/null
@@ -1,9 +0,0 @@
-import os
-from .clip_encoder import CLIPVisionTower
-
-def build_vision_tower(vision_tower_cfg, **kwargs):
- vision_tower = getattr(vision_tower_cfg, 'mm_vision_tower', getattr(vision_tower_cfg, 'vision_tower', None))
- is_absolute_path_exists = os.path.exists(vision_tower)
- if is_absolute_path_exists or vision_tower.startswith("openai") or vision_tower.startswith("laion"):
- return CLIPVisionTower(vision_tower, args=vision_tower_cfg, **kwargs)
-
diff --git a/LLaVA-Med/llava/model/multimodal_encoder/clip_encoder.py b/LLaVA-Med/llava/model/multimodal_encoder/clip_encoder.py
deleted file mode 100644
index dbb9015b0fc9fa93483ba77cc303b793e86c36fc..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/model/multimodal_encoder/clip_encoder.py
+++ /dev/null
@@ -1,78 +0,0 @@
-import torch
-import torch.nn as nn
-
-from transformers import CLIPVisionModel, CLIPImageProcessor, CLIPVisionConfig
-
-
-class CLIPVisionTower(nn.Module):
- def __init__(self, vision_tower, args, delay_load=False):
- super().__init__()
-
- self.is_loaded = False
-
- self.vision_tower_name = vision_tower
- self.select_layer = args.mm_vision_select_layer
- self.select_feature = getattr(args, 'mm_vision_select_feature', 'patch')
-
- if not delay_load:
- self.load_model()
- else:
- self.cfg_only = CLIPVisionConfig.from_pretrained(self.vision_tower_name)
-
- def load_model(self):
- self.image_processor = CLIPImageProcessor.from_pretrained(self.vision_tower_name)
- self.vision_tower = CLIPVisionModel.from_pretrained(self.vision_tower_name)
- self.vision_tower.requires_grad_(False)
-
- self.is_loaded = True
-
- def feature_select(self, image_forward_outs):
- image_features = image_forward_outs.hidden_states[self.select_layer]
- if self.select_feature == 'patch':
- image_features = image_features[:, 1:]
- elif self.select_feature == 'cls_patch':
- image_features = image_features
- else:
- raise ValueError(f'Unexpected select feature: {self.select_feature}')
- return image_features
-
- @torch.no_grad()
- def forward(self, images):
- if type(images) is list:
- image_features = []
- for image in images:
- image_forward_out = self.vision_tower(image.to(device=self.device, dtype=self.dtype).unsqueeze(0), output_hidden_states=True)
- image_feature = self.feature_select(image_forward_out).to(image.dtype)
- image_features.append(image_feature)
- else:
- image_forward_outs = self.vision_tower(images.to(device=self.device, dtype=self.dtype), output_hidden_states=True)
- image_features = self.feature_select(image_forward_outs).to(images.dtype)
-
- return image_features
-
- @property
- def dummy_feature(self):
- return torch.zeros(1, self.hidden_size, device=self.device, dtype=self.dtype)
-
- @property
- def dtype(self):
- return self.vision_tower.dtype
-
- @property
- def device(self):
- return self.vision_tower.device
-
- @property
- def config(self):
- if self.is_loaded:
- return self.vision_tower.config
- else:
- return self.cfg_only
-
- @property
- def hidden_size(self):
- return self.config.hidden_size
-
- @property
- def num_patches(self):
- return (self.config.image_size // self.config.patch_size) ** 2
diff --git a/LLaVA-Med/llava/model/multimodal_projector/builder.py b/LLaVA-Med/llava/model/multimodal_projector/builder.py
deleted file mode 100644
index 32bb7255f378e8085a858845da4f1c5daa132dc1..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/model/multimodal_projector/builder.py
+++ /dev/null
@@ -1,51 +0,0 @@
-import torch
-import torch.nn as nn
-import re
-
-
-class IdentityMap(nn.Module):
- def __init__(self):
- super().__init__()
-
- def forward(self, x, *args, **kwargs):
- return x
-
- @property
- def config(self):
- return {"mm_projector_type": 'identity'}
-
-
-class SimpleResBlock(nn.Module):
- def __init__(self, channels):
- super().__init__()
- self.pre_norm = nn.LayerNorm(channels)
-
- self.proj = nn.Sequential(
- nn.Linear(channels, channels),
- nn.GELU(),
- nn.Linear(channels, channels)
- )
- def forward(self, x):
- x = self.pre_norm(x)
- return x + self.proj(x)
-
-
-def build_vision_projector(config, delay_load=False, **kwargs):
- projector_type = getattr(config, 'mm_projector_type', 'linear')
-
- if projector_type == 'linear':
- return nn.Linear(config.mm_hidden_size, config.hidden_size)
-
- mlp_gelu_match = re.match(r'^mlp(\d+)x_gelu$', projector_type)
- if mlp_gelu_match:
- mlp_depth = int(mlp_gelu_match.group(1))
- modules = [nn.Linear(config.mm_hidden_size, config.hidden_size)]
- for _ in range(1, mlp_depth):
- modules.append(nn.GELU())
- modules.append(nn.Linear(config.hidden_size, config.hidden_size))
- return nn.Sequential(*modules)
-
- if projector_type == 'identity':
- return IdentityMap()
-
- raise ValueError(f'Unknown projector type: {projector_type}')
diff --git a/LLaVA-Med/llava/serve/__init__.py b/LLaVA-Med/llava/serve/__init__.py
deleted file mode 100644
index e69de29bb2d1d6434b8b29ae775ad8c2e48c5391..0000000000000000000000000000000000000000
diff --git a/LLaVA-Med/llava/serve/cli.py b/LLaVA-Med/llava/serve/cli.py
deleted file mode 100644
index 7c73f8869235e63b31c43a0a4071903a45dff0b5..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/serve/cli.py
+++ /dev/null
@@ -1,125 +0,0 @@
-import argparse
-import torch
-
-from llava.constants import IMAGE_TOKEN_INDEX, DEFAULT_IMAGE_TOKEN, DEFAULT_IM_START_TOKEN, DEFAULT_IM_END_TOKEN
-from llava.conversation import conv_templates, SeparatorStyle
-from llava.model.builder import load_pretrained_model
-from llava.utils import disable_torch_init
-from llava.mm_utils import process_images, tokenizer_image_token, get_model_name_from_path, KeywordsStoppingCriteria
-
-from PIL import Image
-
-import requests
-from PIL import Image
-from io import BytesIO
-from transformers import TextStreamer
-
-
-def load_image(image_file):
- if image_file.startswith('http://') or image_file.startswith('https://'):
- response = requests.get(image_file)
- image = Image.open(BytesIO(response.content)).convert('RGB')
- else:
- image = Image.open(image_file).convert('RGB')
- return image
-
-
-def main(args):
- # Model
- disable_torch_init()
-
- model_name = get_model_name_from_path(args.model_path)
- tokenizer, model, image_processor, context_len = load_pretrained_model(args.model_path, args.model_base, model_name, args.load_8bit, args.load_4bit, device=args.device)
-
- if 'llama-2' in model_name.lower():
- conv_mode = "llava_llama_2"
- elif "v1" in model_name.lower():
- conv_mode = "llava_v1"
- elif "mpt" in model_name.lower():
- conv_mode = "mpt"
- else:
- conv_mode = "llava_v0"
- conv_mode = "mistral_instruct"
-
- if args.conv_mode is not None and conv_mode != args.conv_mode:
- print('[WARNING] the auto inferred conversation mode is {}, while `--conv-mode` is {}, using {}'.format(conv_mode, args.conv_mode, args.conv_mode))
- else:
- args.conv_mode = conv_mode
-
- conv = conv_templates[args.conv_mode].copy()
- if "mpt" in model_name.lower():
- roles = ('user', 'assistant')
- else:
- roles = conv.roles
-
- image = load_image(args.image_file)
- # Similar operation in model_worker.py
- image_tensor = process_images([image], image_processor, model.config)
- if type(image_tensor) is list:
- image_tensor = [image.to(model.device, dtype=torch.float16) for image in image_tensor]
- else:
- image_tensor = image_tensor.to(model.device, dtype=torch.float16)
-
- while True:
- try:
- inp = input(f"{roles[0]}: ")
- except EOFError:
- inp = ""
- if not inp:
- print("exit...")
- break
-
- print(f"{roles[1]}: ", end="")
-
- if image is not None:
- # first message
- if model.config.mm_use_im_start_end:
- inp = DEFAULT_IM_START_TOKEN + DEFAULT_IMAGE_TOKEN + DEFAULT_IM_END_TOKEN + '\n' + inp
- else:
- inp = DEFAULT_IMAGE_TOKEN + '\n' + inp
- conv.append_message(conv.roles[0], inp)
- image = None
- else:
- # later messages
- conv.append_message(conv.roles[0], inp)
- conv.append_message(conv.roles[1], None)
- prompt = conv.get_prompt()
-
- input_ids = tokenizer_image_token(prompt, tokenizer, IMAGE_TOKEN_INDEX, return_tensors='pt').unsqueeze(0).to(model.device)
- stop_str = conv.sep if conv.sep_style != SeparatorStyle.TWO else conv.sep2
- keywords = [stop_str]
- stopping_criteria = KeywordsStoppingCriteria(keywords, tokenizer, input_ids)
- streamer = TextStreamer(tokenizer, skip_prompt=True, skip_special_tokens=True)
-
- with torch.inference_mode():
- output_ids = model.generate(
- input_ids,
- images=image_tensor,
- do_sample=True if args.temperature > 0 else False,
- temperature=args.temperature,
- max_new_tokens=args.max_new_tokens,
- streamer=streamer,
- use_cache=True,
- stopping_criteria=[stopping_criteria])
-
- outputs = tokenizer.decode(output_ids[0, input_ids.shape[1]:]).strip()
- conv.messages[-1][-1] = outputs
-
- if args.debug:
- print("\n", {"prompt": prompt, "outputs": outputs}, "\n")
-
-
-if __name__ == "__main__":
- parser = argparse.ArgumentParser()
- parser.add_argument("--model-path", type=str, default="facebook/opt-350m")
- parser.add_argument("--model-base", type=str, default=None)
- parser.add_argument("--image-file", type=str, required=True)
- parser.add_argument("--device", type=str, default="cuda")
- parser.add_argument("--conv-mode", type=str, default=None)
- parser.add_argument("--temperature", type=float, default=0.2)
- parser.add_argument("--max-new-tokens", type=int, default=512)
- parser.add_argument("--load-8bit", action="store_true")
- parser.add_argument("--load-4bit", action="store_true")
- parser.add_argument("--debug", action="store_true")
- args = parser.parse_args()
- main(args)
diff --git a/LLaVA-Med/llava/serve/controller.py b/LLaVA-Med/llava/serve/controller.py
deleted file mode 100644
index b61fca6ea9fe8aa37acd143784a3d76e90a58b9f..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/serve/controller.py
+++ /dev/null
@@ -1,298 +0,0 @@
-"""
-A controller manages distributed workers.
-It sends worker addresses to clients.
-"""
-import argparse
-import asyncio
-import dataclasses
-from enum import Enum, auto
-import json
-import logging
-import time
-from typing import List, Union
-import threading
-
-from fastapi import FastAPI, Request
-from fastapi.responses import StreamingResponse
-import numpy as np
-import requests
-import uvicorn
-
-from llava.constants import CONTROLLER_HEART_BEAT_EXPIRATION
-from llava.utils import build_logger, server_error_msg
-
-
-logger = build_logger("controller", "controller.log")
-
-
-class DispatchMethod(Enum):
- LOTTERY = auto()
- SHORTEST_QUEUE = auto()
-
- @classmethod
- def from_str(cls, name):
- if name == "lottery":
- return cls.LOTTERY
- elif name == "shortest_queue":
- return cls.SHORTEST_QUEUE
- else:
- raise ValueError(f"Invalid dispatch method")
-
-
-@dataclasses.dataclass
-class WorkerInfo:
- model_names: List[str]
- speed: int
- queue_length: int
- check_heart_beat: bool
- last_heart_beat: str
-
-
-def heart_beat_controller(controller):
- while True:
- time.sleep(CONTROLLER_HEART_BEAT_EXPIRATION)
- controller.remove_stable_workers_by_expiration()
-
-
-class Controller:
- def __init__(self, dispatch_method: str):
- # Dict[str -> WorkerInfo]
- self.worker_info = {}
- self.dispatch_method = DispatchMethod.from_str(dispatch_method)
-
- self.heart_beat_thread = threading.Thread(
- target=heart_beat_controller, args=(self,))
- self.heart_beat_thread.start()
-
- logger.info("Init controller")
-
- def register_worker(self, worker_name: str, check_heart_beat: bool,
- worker_status: dict):
- if worker_name not in self.worker_info:
- logger.info(f"Register a new worker: {worker_name}")
- else:
- logger.info(f"Register an existing worker: {worker_name}")
-
- if not worker_status:
- worker_status = self.get_worker_status(worker_name)
- if not worker_status:
- return False
-
- self.worker_info[worker_name] = WorkerInfo(
- worker_status["model_names"], worker_status["speed"], worker_status["queue_length"],
- check_heart_beat, time.time())
-
- logger.info(f"Register done: {worker_name}, {worker_status}")
- return True
-
- def get_worker_status(self, worker_name: str):
- try:
- r = requests.post(worker_name + "/worker_get_status", timeout=5)
- except requests.exceptions.RequestException as e:
- logger.error(f"Get status fails: {worker_name}, {e}")
- return None
-
- if r.status_code != 200:
- logger.error(f"Get status fails: {worker_name}, {r}")
- return None
-
- return r.json()
-
- def remove_worker(self, worker_name: str):
- del self.worker_info[worker_name]
-
- def refresh_all_workers(self):
- old_info = dict(self.worker_info)
- self.worker_info = {}
-
- for w_name, w_info in old_info.items():
- if not self.register_worker(w_name, w_info.check_heart_beat, None):
- logger.info(f"Remove stale worker: {w_name}")
-
- def list_models(self):
- model_names = set()
-
- for w_name, w_info in self.worker_info.items():
- model_names.update(w_info.model_names)
-
- return list(model_names)
-
- def get_worker_address(self, model_name: str):
- if self.dispatch_method == DispatchMethod.LOTTERY:
- worker_names = []
- worker_speeds = []
- for w_name, w_info in self.worker_info.items():
- if model_name in w_info.model_names:
- worker_names.append(w_name)
- worker_speeds.append(w_info.speed)
- worker_speeds = np.array(worker_speeds, dtype=np.float32)
- norm = np.sum(worker_speeds)
- if norm < 1e-4:
- return ""
- worker_speeds = worker_speeds / norm
- if True: # Directly return address
- pt = np.random.choice(np.arange(len(worker_names)),
- p=worker_speeds)
- worker_name = worker_names[pt]
- return worker_name
-
- # Check status before returning
- while True:
- pt = np.random.choice(np.arange(len(worker_names)),
- p=worker_speeds)
- worker_name = worker_names[pt]
-
- if self.get_worker_status(worker_name):
- break
- else:
- self.remove_worker(worker_name)
- worker_speeds[pt] = 0
- norm = np.sum(worker_speeds)
- if norm < 1e-4:
- return ""
- worker_speeds = worker_speeds / norm
- continue
- return worker_name
- elif self.dispatch_method == DispatchMethod.SHORTEST_QUEUE:
- worker_names = []
- worker_qlen = []
- for w_name, w_info in self.worker_info.items():
- if model_name in w_info.model_names:
- worker_names.append(w_name)
- worker_qlen.append(w_info.queue_length / w_info.speed)
- if len(worker_names) == 0:
- return ""
- min_index = np.argmin(worker_qlen)
- w_name = worker_names[min_index]
- self.worker_info[w_name].queue_length += 1
- logger.info(f"names: {worker_names}, queue_lens: {worker_qlen}, ret: {w_name}")
- return w_name
- else:
- raise ValueError(f"Invalid dispatch method: {self.dispatch_method}")
-
- def receive_heart_beat(self, worker_name: str, queue_length: int):
- if worker_name not in self.worker_info:
- logger.info(f"Receive unknown heart beat. {worker_name}")
- return False
-
- self.worker_info[worker_name].queue_length = queue_length
- self.worker_info[worker_name].last_heart_beat = time.time()
- logger.info(f"Receive heart beat. {worker_name}")
- return True
-
- def remove_stable_workers_by_expiration(self):
- expire = time.time() - CONTROLLER_HEART_BEAT_EXPIRATION
- to_delete = []
- for worker_name, w_info in self.worker_info.items():
- if w_info.check_heart_beat and w_info.last_heart_beat < expire:
- to_delete.append(worker_name)
-
- for worker_name in to_delete:
- self.remove_worker(worker_name)
-
- def worker_api_generate_stream(self, params):
- worker_addr = self.get_worker_address(params["model"])
- if not worker_addr:
- logger.info(f"no worker: {params['model']}")
- ret = {
- "text": server_error_msg,
- "error_code": 2,
- }
- yield json.dumps(ret).encode() + b"\0"
-
- try:
- response = requests.post(worker_addr + "/worker_generate_stream",
- json=params, stream=True, timeout=5)
- for chunk in response.iter_lines(decode_unicode=False, delimiter=b"\0"):
- if chunk:
- yield chunk + b"\0"
- except requests.exceptions.RequestException as e:
- logger.info(f"worker timeout: {worker_addr}")
- ret = {
- "text": server_error_msg,
- "error_code": 3,
- }
- yield json.dumps(ret).encode() + b"\0"
-
-
- # Let the controller act as a worker to achieve hierarchical
- # management. This can be used to connect isolated sub networks.
- def worker_api_get_status(self):
- model_names = set()
- speed = 0
- queue_length = 0
-
- for w_name in self.worker_info:
- worker_status = self.get_worker_status(w_name)
- if worker_status is not None:
- model_names.update(worker_status["model_names"])
- speed += worker_status["speed"]
- queue_length += worker_status["queue_length"]
-
- return {
- "model_names": list(model_names),
- "speed": speed,
- "queue_length": queue_length,
- }
-
-
-app = FastAPI()
-
-
-@app.post("/register_worker")
-async def register_worker(request: Request):
- data = await request.json()
- controller.register_worker(
- data["worker_name"], data["check_heart_beat"],
- data.get("worker_status", None))
-
-
-@app.post("/refresh_all_workers")
-async def refresh_all_workers():
- models = controller.refresh_all_workers()
-
-
-@app.post("/list_models")
-async def list_models():
- models = controller.list_models()
- return {"models": models}
-
-
-@app.post("/get_worker_address")
-async def get_worker_address(request: Request):
- data = await request.json()
- addr = controller.get_worker_address(data["model"])
- return {"address": addr}
-
-
-@app.post("/receive_heart_beat")
-async def receive_heart_beat(request: Request):
- data = await request.json()
- exist = controller.receive_heart_beat(
- data["worker_name"], data["queue_length"])
- return {"exist": exist}
-
-
-@app.post("/worker_generate_stream")
-async def worker_api_generate_stream(request: Request):
- params = await request.json()
- generator = controller.worker_api_generate_stream(params)
- return StreamingResponse(generator)
-
-
-@app.post("/worker_get_status")
-async def worker_api_get_status(request: Request):
- return controller.worker_api_get_status()
-
-
-if __name__ == "__main__":
- parser = argparse.ArgumentParser()
- parser.add_argument("--host", type=str, default="localhost")
- parser.add_argument("--port", type=int, default=21001)
- parser.add_argument("--dispatch-method", type=str, choices=[
- "lottery", "shortest_queue"], default="shortest_queue")
- args = parser.parse_args()
- logger.info(f"args: {args}")
-
- controller = Controller(args.dispatch_method)
- uvicorn.run(app, host=args.host, port=args.port, log_level="info")
diff --git a/LLaVA-Med/llava/serve/examples/bio_patch.png b/LLaVA-Med/llava/serve/examples/bio_patch.png
deleted file mode 100644
index 26ad078703c198e7ea03f25f63d191d34174eb29..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/llava/serve/examples/bio_patch.png and /dev/null differ
diff --git a/LLaVA-Med/llava/serve/examples/extreme_ironing.jpg b/LLaVA-Med/llava/serve/examples/extreme_ironing.jpg
deleted file mode 100644
index 638b078837f175039b2db49a63821288d9681daa..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/llava/serve/examples/extreme_ironing.jpg and /dev/null differ
diff --git a/LLaVA-Med/llava/serve/examples/med_img_1.png b/LLaVA-Med/llava/serve/examples/med_img_1.png
deleted file mode 100644
index 3d3832ac82dd29296986d723e4dbe24512024770..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/llava/serve/examples/med_img_1.png and /dev/null differ
diff --git a/LLaVA-Med/llava/serve/examples/synpic32933.jpg b/LLaVA-Med/llava/serve/examples/synpic32933.jpg
deleted file mode 100644
index c85669739e3237b9020366386b52beba5807a1d2..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/llava/serve/examples/synpic32933.jpg and /dev/null differ
diff --git a/LLaVA-Med/llava/serve/examples/synpic42202.jpg b/LLaVA-Med/llava/serve/examples/synpic42202.jpg
deleted file mode 100644
index 3fdfd4dbb33ca2db39c66fb8276c6ee3c203f8de..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/llava/serve/examples/synpic42202.jpg and /dev/null differ
diff --git a/LLaVA-Med/llava/serve/examples/waterview.jpg b/LLaVA-Med/llava/serve/examples/waterview.jpg
deleted file mode 100644
index 6f44ebaba1aa493b8bab3baa4e827b76752b1869..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/llava/serve/examples/waterview.jpg and /dev/null differ
diff --git a/LLaVA-Med/llava/serve/examples/xy_chromosome.jpg b/LLaVA-Med/llava/serve/examples/xy_chromosome.jpg
deleted file mode 100644
index 6e107439858250c85249b05ff16b435cf4f5d03c..0000000000000000000000000000000000000000
Binary files a/LLaVA-Med/llava/serve/examples/xy_chromosome.jpg and /dev/null differ
diff --git a/LLaVA-Med/llava/serve/gradio_web_server.py b/LLaVA-Med/llava/serve/gradio_web_server.py
deleted file mode 100644
index 6f3cf9aa0769306535ffdd2133c1b6c6c5b3234a..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/serve/gradio_web_server.py
+++ /dev/null
@@ -1,477 +0,0 @@
-import argparse
-import datetime
-import json
-import os
-import time
-
-import gradio as gr
-import requests
-
-from llava.conversation import (default_conversation, conv_templates,
- SeparatorStyle)
-from llava.constants import LOGDIR
-from llava.utils import (build_logger, server_error_msg,
- violates_moderation, moderation_msg)
-import hashlib
-
-
-logger = build_logger("gradio_web_server", "gradio_web_server.log")
-
-headers = {"User-Agent": "LLaVA-Med Client"}
-
-no_change_btn = gr.Button.update()
-enable_btn = gr.Button.update(interactive=True)
-disable_btn = gr.Button.update(interactive=False)
-
-priority = {
- "vicuna-13b": "aaaaaaa",
- "koala-13b": "aaaaaab",
-}
-
-
-def get_conv_log_filename():
- t = datetime.datetime.now()
- name = os.path.join(LOGDIR, f"{t.year}-{t.month:02d}-{t.day:02d}-conv.json")
- return name
-
-
-def get_model_list():
- ret = requests.post(args.controller_url + "/refresh_all_workers")
- assert ret.status_code == 200
- ret = requests.post(args.controller_url + "/list_models")
- models = ret.json()["models"]
- models.sort(key=lambda x: priority.get(x, x))
- logger.info(f"Models: {models}")
- return models
-
-
-get_window_url_params = """
-function() {
- const params = new URLSearchParams(window.location.search);
- url_params = Object.fromEntries(params);
- console.log(url_params);
- return url_params;
- }
-"""
-
-
-def load_demo(url_params, request: gr.Request):
- logger.info(f"load_demo. ip: {request.client.host}. params: {url_params}")
-
- dropdown_update = gr.Dropdown.update(visible=True)
- if "model" in url_params:
- model = url_params["model"]
- if model in models:
- dropdown_update = gr.Dropdown.update(
- value=model, visible=True)
-
- state = default_conversation.copy()
- return state, dropdown_update
-
-
-def load_demo_refresh_model_list(request: gr.Request):
- logger.info(f"load_demo. ip: {request.client.host}")
- models = get_model_list()
- state = default_conversation.copy()
- dropdown_update = gr.Dropdown.update(
- choices=models,
- value=models[0] if len(models) > 0 else ""
- )
- return state, dropdown_update
-
-
-def vote_last_response(state, vote_type, model_selector, request: gr.Request):
- with open(get_conv_log_filename(), "a") as fout:
- data = {
- "tstamp": round(time.time(), 4),
- "type": vote_type,
- "model": model_selector,
- "state": state.dict(),
- "ip": request.client.host,
- }
- fout.write(json.dumps(data) + "\n")
-
-
-def upvote_last_response(state, model_selector, request: gr.Request):
- logger.info(f"upvote. ip: {request.client.host}")
- vote_last_response(state, "upvote", model_selector, request)
- return ("",) + (disable_btn,) * 3
-
-
-def downvote_last_response(state, model_selector, request: gr.Request):
- logger.info(f"downvote. ip: {request.client.host}")
- vote_last_response(state, "downvote", model_selector, request)
- return ("",) + (disable_btn,) * 3
-
-
-def flag_last_response(state, model_selector, request: gr.Request):
- logger.info(f"flag. ip: {request.client.host}")
- vote_last_response(state, "flag", model_selector, request)
- return ("",) + (disable_btn,) * 3
-
-
-def regenerate(state, image_process_mode, request: gr.Request):
- logger.info(f"regenerate. ip: {request.client.host}")
- state.messages[-1][-1] = None
- prev_human_msg = state.messages[-2]
- if type(prev_human_msg[1]) in (tuple, list):
- prev_human_msg[1] = (*prev_human_msg[1][:2], image_process_mode)
- state.skip_next = False
- return (state, state.to_gradio_chatbot(), "", None) + (disable_btn,) * 5
-
-
-def clear_history(request: gr.Request):
- logger.info(f"clear_history. ip: {request.client.host}")
- state = default_conversation.copy()
- return (state, state.to_gradio_chatbot(), "", None) + (disable_btn,) * 5
-
-
-def add_text(state, text, image, image_process_mode, request: gr.Request):
- logger.info(f"add_text. ip: {request.client.host}. len: {len(text)}")
- if len(text) <= 0 and image is None:
- state.skip_next = True
- return (state, state.to_gradio_chatbot(), "", None) + (no_change_btn,) * 5
- if args.moderate:
- flagged = violates_moderation(text)
- if flagged:
- state.skip_next = True
- return (state, state.to_gradio_chatbot(), moderation_msg, None) + (
- no_change_btn,) * 5
-
- text = text[:1536] # Hard cut-off
- if image is not None:
- text = text[:1200] # Hard cut-off for images
- if '' not in text:
- # text = '' + text
- text = text + '\n'
- text = (text, image, image_process_mode)
- if len(state.get_images(return_pil=True)) > 0:
- state = default_conversation.copy()
- state.append_message(state.roles[0], text)
- state.append_message(state.roles[1], None)
- state.skip_next = False
- return (state, state.to_gradio_chatbot(), "", None) + (disable_btn,) * 5
-
-
-def http_bot(state, model_selector, temperature, top_p, max_new_tokens, request: gr.Request):
- logger.info(f"http_bot. ip: {request.client.host}")
- start_tstamp = time.time()
- model_name = model_selector
-
- if state.skip_next:
- # This generate call is skipped due to invalid inputs
- yield (state, state.to_gradio_chatbot()) + (no_change_btn,) * 5
- return
-
- if len(state.messages) == state.offset + 2:
- # First round of conversation
- if "llava" in model_name.lower():
- if 'llama-2' in model_name.lower():
- template_name = "llava_llama_2"
- elif "v1" in model_name.lower():
- if 'mmtag' in model_name.lower():
- template_name = "v1_mmtag"
- elif 'plain' in model_name.lower() and 'finetune' not in model_name.lower():
- template_name = "v1_mmtag"
- else:
- template_name = "llava_v1"
- elif "mpt" in model_name.lower():
- template_name = "mpt"
- else:
- if 'mmtag' in model_name.lower():
- template_name = "v0_mmtag"
- elif 'plain' in model_name.lower() and 'finetune' not in model_name.lower():
- template_name = "v0_mmtag"
- else:
- template_name = "llava_v0"
- elif "mpt" in model_name:
- template_name = "mpt_text"
- elif "llama-2" in model_name:
- template_name = "llama_2"
- else:
- template_name = "vicuna_v1"
- template_name = "mistral_instruct" # FIXME: overwrite
- new_state = conv_templates[template_name].copy()
- new_state.append_message(new_state.roles[0], state.messages[-2][1])
- new_state.append_message(new_state.roles[1], None)
- state = new_state
-
- # Query worker address
- controller_url = args.controller_url
- ret = requests.post(controller_url + "/get_worker_address",
- json={"model": model_name})
- worker_addr = ret.json()["address"]
- logger.info(f"model_name: {model_name}, worker_addr: {worker_addr}")
-
- # No available worker
- if worker_addr == "":
- state.messages[-1][-1] = server_error_msg
- yield (state, state.to_gradio_chatbot(), disable_btn, disable_btn, disable_btn, enable_btn, enable_btn)
- return
-
- # Construct prompt
- prompt = state.get_prompt()
-
- all_images = state.get_images(return_pil=True)
- all_image_hash = [hashlib.md5(image.tobytes()).hexdigest() for image in all_images]
- for image, hash in zip(all_images, all_image_hash):
- t = datetime.datetime.now()
- filename = os.path.join(LOGDIR, "serve_images", f"{t.year}-{t.month:02d}-{t.day:02d}", f"{hash}.jpg")
- if not os.path.isfile(filename):
- os.makedirs(os.path.dirname(filename), exist_ok=True)
- image.save(filename)
-
- # Make requests
- pload = {
- "model": model_name,
- "prompt": prompt,
- "temperature": float(temperature),
- "top_p": float(top_p),
- "max_new_tokens": min(int(max_new_tokens), 1536),
- "stop": state.sep if state.sep_style in [SeparatorStyle.SINGLE, SeparatorStyle.MPT] else state.sep2,
- "images": f'List of {len(state.get_images())} images: {all_image_hash}',
- }
- logger.info(f"==== request ====\n{pload}")
-
- pload['images'] = state.get_images()
-
- state.messages[-1][-1] = "▌"
- yield (state, state.to_gradio_chatbot()) + (disable_btn,) * 5
-
- try:
- # Stream output
- response = requests.post(worker_addr + "/worker_generate_stream",
- headers=headers, json=pload, stream=True, timeout=10)
- for chunk in response.iter_lines(decode_unicode=False, delimiter=b"\0"):
- if chunk:
- data = json.loads(chunk.decode())
- if data["error_code"] == 0:
- output = data["text"][len(prompt):].strip()
- state.messages[-1][-1] = output + "▌"
- yield (state, state.to_gradio_chatbot()) + (disable_btn,) * 5
- else:
- output = data["text"] + f" (error_code: {data['error_code']})"
- state.messages[-1][-1] = output
- yield (state, state.to_gradio_chatbot()) + (disable_btn, disable_btn, disable_btn, enable_btn, enable_btn)
- return
- time.sleep(0.03)
- except requests.exceptions.RequestException as e:
- state.messages[-1][-1] = server_error_msg
- yield (state, state.to_gradio_chatbot()) + (disable_btn, disable_btn, disable_btn, enable_btn, enable_btn)
- return
-
- state.messages[-1][-1] = state.messages[-1][-1][:-1]
- yield (state, state.to_gradio_chatbot()) + (enable_btn,) * 5
-
- finish_tstamp = time.time()
- logger.info(f"{output}")
-
- with open(get_conv_log_filename(), "a") as fout:
- data = {
- "tstamp": round(finish_tstamp, 4),
- "type": "chat",
- "model": model_name,
- "start": round(start_tstamp, 4),
- "finish": round(finish_tstamp, 4),
- "state": state.dict(),
- "images": all_image_hash,
- "ip": request.client.host,
- }
- fout.write(json.dumps(data) + "\n")
-
-
-title_markdown = ("""
-# 🌋 LLaVA-Med: Large Language and Vision Assistant for Medical Research
-[[Project Page]](https://llava-vl.github.io) [[Paper]](https://arxiv.org/abs/2304.08485) [[Code]](https://github.com/haotian-liu/LLaVA) [[Model]](https://huggingface.co/liuhaotian/LLaVA-13b-delta-v0)
-""")
-
-tos_markdown = ("""
-### Terms of use
-By using this service, users are required to agree to the following terms:
-The service is a research preview intended for non-commercial use only. It only provides limited safety measures and may generate offensive content. It must not be used for any illegal, harmful, violent, racist, or sexual purposes. The service may collect user dialogue data for future research.
-Please click the "Flag" button if you get any inappropriate answer! We will collect those to keep improving our moderator.
-For an optimal experience, please use desktop computers for this demo, as mobile devices may compromise its quality.
-""")
-
-
-learn_more_markdown = ("""
-### License
-The service is a research preview intended for non-commercial use only, subject to the model [License](https://github.com/facebookresearch/llama/blob/main/MODEL_CARD.md) of LLaMA, [Terms of Use](https://openai.com/policies/terms-of-use) of the data generated by OpenAI, and [Privacy Practices](https://chrome.google.com/webstore/detail/sharegpt-share-your-chatg/daiacboceoaocpibfodeljbdfacokfjb) of ShareGPT. Please contact us if you find any potential violation.
-""")
-
-block_css = """
-
-#buttons button {
- min-width: min(120px,100%);
-}
-
-"""
-
-def build_demo(embed_mode):
- textbox = gr.Textbox(show_label=False, placeholder="Enter text and press ENTER", container=False)
- with gr.Blocks(title="LLaVA", theme=gr.themes.Default(), css=block_css) as demo:
- state = gr.State()
-
- if not embed_mode:
- gr.Markdown(title_markdown)
-
- with gr.Row():
- with gr.Column(scale=3):
- with gr.Row(elem_id="model_selector_row"):
- model_selector = gr.Dropdown(
- choices=models,
- value=models[0] if len(models) > 0 else "",
- interactive=True,
- show_label=False,
- container=False)
-
- imagebox = gr.Image(type="pil")
- image_process_mode = gr.Radio(
- ["Crop", "Resize", "Pad", "Default"],
- value="Default",
- label="Preprocess for non-square image", visible=False)
-
- cur_dir = os.path.dirname(os.path.abspath(__file__))
- gr.Examples(examples=[
- [f"{cur_dir}/examples/bio_patch.png", "What is this image about?"],
- [f"{cur_dir}/examples/med_img_1.png", "Can you describe the image in details?"],
- [f"{cur_dir}/examples/xy_chromosome.jpg", "Can you describe the image in details?"],
- [f"{cur_dir}/examples/synpic42202.jpg", "Is there evidence of an aortic aneurysm? Please choose from the following two options: [yes, no]?"], # answer" yes
- [f"{cur_dir}/examples/synpic32933.jpg", "What is the abnormality by the right hemidiaphragm?"], # answer: free air
- [f"{cur_dir}/examples/extreme_ironing.jpg", "What is unusual about this image?"],
- [f"{cur_dir}/examples/waterview.jpg", "What are the things I should be cautious about when I visit here?"],
- ], inputs=[imagebox, textbox])
-
- with gr.Accordion("Parameters", open=False) as parameter_row:
- temperature = gr.Slider(minimum=0.0, maximum=1.0, value=0.2, step=0.1, interactive=True, label="Temperature",)
- top_p = gr.Slider(minimum=0.0, maximum=1.0, value=0.7, step=0.1, interactive=True, label="Top P",)
- max_output_tokens = gr.Slider(minimum=0, maximum=1024, value=512, step=64, interactive=True, label="Max output tokens",)
-
- with gr.Column(scale=8):
- chatbot = gr.Chatbot(elem_id="chatbot", label="LLaVA-Med Chatbot", height=550)
- with gr.Row():
- with gr.Column(scale=8):
- textbox.render()
- with gr.Column(scale=1, min_width=50):
- submit_btn = gr.Button(value="Send", variant="primary")
- with gr.Row(elem_id="buttons") as button_row:
- upvote_btn = gr.Button(value="👍 Upvote", interactive=False)
- downvote_btn = gr.Button(value="👎 Downvote", interactive=False)
- flag_btn = gr.Button(value="⚠️ Flag", interactive=False)
- #stop_btn = gr.Button(value="⏹️ Stop Generation", interactive=False)
- regenerate_btn = gr.Button(value="🔄 Regenerate", interactive=False)
- clear_btn = gr.Button(value="🗑️ Clear", interactive=False)
-
- if not embed_mode:
- gr.Markdown(tos_markdown)
- gr.Markdown(learn_more_markdown)
- url_params = gr.JSON(visible=False)
-
- # Register listeners
- btn_list = [upvote_btn, downvote_btn, flag_btn, regenerate_btn, clear_btn]
- upvote_btn.click(
- upvote_last_response,
- [state, model_selector],
- [textbox, upvote_btn, downvote_btn, flag_btn],
- queue=False
- )
- downvote_btn.click(
- downvote_last_response,
- [state, model_selector],
- [textbox, upvote_btn, downvote_btn, flag_btn],
- queue=False
- )
- flag_btn.click(
- flag_last_response,
- [state, model_selector],
- [textbox, upvote_btn, downvote_btn, flag_btn],
- queue=False
- )
-
- regenerate_btn.click(
- regenerate,
- [state, image_process_mode],
- [state, chatbot, textbox, imagebox] + btn_list,
- queue=False
- ).then(
- http_bot,
- [state, model_selector, temperature, top_p, max_output_tokens],
- [state, chatbot] + btn_list
- )
-
- clear_btn.click(
- clear_history,
- None,
- [state, chatbot, textbox, imagebox] + btn_list,
- queue=False
- )
-
- textbox.submit(
- add_text,
- [state, textbox, imagebox, image_process_mode],
- [state, chatbot, textbox, imagebox] + btn_list,
- queue=False
- ).then(
- http_bot,
- [state, model_selector, temperature, top_p, max_output_tokens],
- [state, chatbot] + btn_list
- )
-
- submit_btn.click(
- add_text,
- [state, textbox, imagebox, image_process_mode],
- [state, chatbot, textbox, imagebox] + btn_list,
- queue=False
- ).then(
- http_bot,
- [state, model_selector, temperature, top_p, max_output_tokens],
- [state, chatbot] + btn_list
- )
-
- if args.model_list_mode == "once":
- demo.load(
- load_demo,
- [url_params],
- [state, model_selector],
- _js=get_window_url_params,
- queue=False
- )
- elif args.model_list_mode == "reload":
- demo.load(
- load_demo_refresh_model_list,
- None,
- [state, model_selector],
- queue=False
- )
- else:
- raise ValueError(f"Unknown model list mode: {args.model_list_mode}")
-
- return demo
-
-
-if __name__ == "__main__":
- parser = argparse.ArgumentParser()
- parser.add_argument("--host", type=str, default="0.0.0.0")
- parser.add_argument("--port", type=int)
- parser.add_argument("--controller-url", type=str, default="http://localhost:21001")
- parser.add_argument("--concurrency-count", type=int, default=10)
- parser.add_argument("--model-list-mode", type=str, default="once",
- choices=["once", "reload"])
- parser.add_argument("--share", action="store_true")
- parser.add_argument("--moderate", action="store_true")
- parser.add_argument("--embed", action="store_true")
- args = parser.parse_args()
- logger.info(f"args: {args}")
-
- models = get_model_list()
-
- logger.info(args)
- demo = build_demo(args.embed)
- demo.queue(
- concurrency_count=args.concurrency_count,
- api_open=False
- ).launch(
- server_name=args.host,
- server_port=args.port,
- share=args.share
- )
diff --git a/LLaVA-Med/llava/serve/model_worker.py b/LLaVA-Med/llava/serve/model_worker.py
deleted file mode 100644
index a7bcd0829d0120c4359400b958fc3ad6c6867f9c..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/serve/model_worker.py
+++ /dev/null
@@ -1,285 +0,0 @@
-"""
-A model worker executes the model.
-"""
-import argparse
-import asyncio
-import json
-import time
-import threading
-import uuid
-
-from fastapi import FastAPI, Request, BackgroundTasks
-from fastapi.responses import StreamingResponse
-import requests
-import torch
-import uvicorn
-from functools import partial
-
-from llava.constants import WORKER_HEART_BEAT_INTERVAL
-from llava.utils import (build_logger, server_error_msg,
- pretty_print_semaphore)
-from llava.model.builder import load_pretrained_model
-from llava.mm_utils import process_images, load_image_from_base64, tokenizer_image_token, KeywordsStoppingCriteria
-from llava.constants import IMAGE_TOKEN_INDEX, DEFAULT_IMAGE_TOKEN, DEFAULT_IM_START_TOKEN, DEFAULT_IM_END_TOKEN
-from transformers import TextIteratorStreamer
-from threading import Thread
-
-
-GB = 1 << 30
-
-worker_id = str(uuid.uuid4())[:6]
-logger = build_logger("model_worker", f"model_worker_{worker_id}.log")
-global_counter = 0
-
-model_semaphore = None
-
-
-def heart_beat_worker(controller):
-
- while True:
- time.sleep(WORKER_HEART_BEAT_INTERVAL)
- controller.send_heart_beat()
-
-
-class ModelWorker:
- def __init__(self, controller_addr, worker_addr,
- worker_id, no_register,
- model_path, model_base, model_name,
- load_8bit, load_4bit, device):
- self.controller_addr = controller_addr
- self.worker_addr = worker_addr
- self.worker_id = worker_id
- if model_path.endswith("/"):
- model_path = model_path[:-1]
- if model_name is None:
- model_paths = model_path.split("/")
- if model_paths[-1].startswith('checkpoint-'):
- self.model_name = model_paths[-2] + "_" + model_paths[-1]
- else:
- self.model_name = model_paths[-1]
- else:
- self.model_name = model_name
-
- self.device = device
- logger.info(f"Loading the model {self.model_name} on worker {worker_id} ...")
- self.tokenizer, self.model, self.image_processor, self.context_len = load_pretrained_model(
- model_path, model_base, self.model_name, load_8bit, load_4bit, device=self.device)
- self.is_multimodal = 'llava' in self.model_name.lower()
-
- if not no_register:
- self.register_to_controller()
- self.heart_beat_thread = threading.Thread(
- target=heart_beat_worker, args=(self,))
- self.heart_beat_thread.start()
-
- def register_to_controller(self):
- logger.info("Register to controller")
-
- url = self.controller_addr + "/register_worker"
- data = {
- "worker_name": self.worker_addr,
- "check_heart_beat": True,
- "worker_status": self.get_status()
- }
- r = requests.post(url, json=data)
- assert r.status_code == 200
-
- def send_heart_beat(self):
- logger.info(f"Send heart beat. Models: {[self.model_name]}. "
- f"Semaphore: {pretty_print_semaphore(model_semaphore)}. "
- f"global_counter: {global_counter}")
-
- url = self.controller_addr + "/receive_heart_beat"
-
- while True:
- try:
- ret = requests.post(url, json={
- "worker_name": self.worker_addr,
- "queue_length": self.get_queue_length()}, timeout=5)
- exist = ret.json()["exist"]
- break
- except requests.exceptions.RequestException as e:
- logger.error(f"heart beat error: {e}")
- time.sleep(5)
-
- if not exist:
- self.register_to_controller()
-
- def get_queue_length(self):
- if model_semaphore is None:
- return 0
- else:
- return args.limit_model_concurrency - model_semaphore._value + (len(
- model_semaphore._waiters) if model_semaphore._waiters is not None else 0)
-
- def get_status(self):
- return {
- "model_names": [self.model_name],
- "speed": 1,
- "queue_length": self.get_queue_length(),
- }
-
- @torch.inference_mode()
- def generate_stream(self, params):
- tokenizer, model, image_processor = self.tokenizer, self.model, self.image_processor
-
- prompt = params["prompt"]
- ori_prompt = prompt
- images = params.get("images", None)
- num_image_tokens = 0
- if images is not None and len(images) > 0 and self.is_multimodal:
- if len(images) > 0:
- if len(images) != prompt.count(DEFAULT_IMAGE_TOKEN):
- raise ValueError("Number of images does not match number of tokens in prompt")
-
- images = [load_image_from_base64(image) for image in images]
- images = process_images(images, image_processor, model.config)
-
- if type(images) is list:
- images = [image.to(self.model.device, dtype=torch.float16) for image in images]
- else:
- images = images.to(self.model.device, dtype=torch.float16)
-
- replace_token = DEFAULT_IMAGE_TOKEN
- if getattr(self.model.config, 'mm_use_im_start_end', False):
- replace_token = DEFAULT_IM_START_TOKEN + replace_token + DEFAULT_IM_END_TOKEN
- prompt = prompt.replace(DEFAULT_IMAGE_TOKEN, replace_token)
-
- num_image_tokens = prompt.count(replace_token) * model.get_vision_tower().num_patches
- else:
- images = None
- image_args = {"images": images}
- else:
- images = None
- image_args = {}
-
- temperature = float(params.get("temperature", 1.0))
- top_p = float(params.get("top_p", 1.0))
- max_context_length = getattr(model.config, 'max_position_embeddings', 2048)
- max_new_tokens = min(int(params.get("max_new_tokens", 256)), 1024)
- stop_str = params.get("stop", None)
- do_sample = True if temperature > 0.001 else False
-
- input_ids = tokenizer_image_token(prompt, tokenizer, IMAGE_TOKEN_INDEX, return_tensors='pt').unsqueeze(0).to(self.device)
- keywords = [stop_str]
- stopping_criteria = KeywordsStoppingCriteria(keywords, tokenizer, input_ids)
- streamer = TextIteratorStreamer(tokenizer, skip_prompt=True, skip_special_tokens=True, timeout=15)
-
- max_new_tokens = min(max_new_tokens, max_context_length - input_ids.shape[-1] - num_image_tokens)
-
- if max_new_tokens < 1:
- yield json.dumps({"text": ori_prompt + "Exceeds max token length. Please start a new conversation, thanks.", "error_code": 0}).encode() + b"\0"
- return
-
- thread = Thread(target=model.generate, kwargs=dict(
- inputs=input_ids,
- do_sample=do_sample,
- temperature=temperature,
- top_p=top_p,
- max_new_tokens=max_new_tokens,
- streamer=streamer,
- stopping_criteria=[stopping_criteria],
- use_cache=True,
- **image_args
- ))
- thread.start()
-
- generated_text = ori_prompt
- for new_text in streamer:
- generated_text += new_text
- if generated_text.endswith(stop_str):
- generated_text = generated_text[:-len(stop_str)]
- yield json.dumps({"text": generated_text, "error_code": 0}).encode() + b"\0"
-
- def generate_stream_gate(self, params):
- try:
- for x in self.generate_stream(params):
- yield x
- except ValueError as e:
- print("Caught ValueError:", e)
- ret = {
- "text": server_error_msg,
- "error_code": 1,
- }
- yield json.dumps(ret).encode() + b"\0"
- except torch.cuda.CudaError as e:
- print("Caught torch.cuda.CudaError:", e)
- ret = {
- "text": server_error_msg,
- "error_code": 1,
- }
- yield json.dumps(ret).encode() + b"\0"
- except Exception as e:
- print("Caught Unknown Error", e)
- ret = {
- "text": server_error_msg,
- "error_code": 1,
- }
- yield json.dumps(ret).encode() + b"\0"
-
-
-app = FastAPI()
-
-
-def release_model_semaphore(fn=None):
- model_semaphore.release()
- if fn is not None:
- fn()
-
-
-@app.post("/worker_generate_stream")
-async def generate_stream(request: Request):
- global model_semaphore, global_counter
- global_counter += 1
- params = await request.json()
-
- if model_semaphore is None:
- model_semaphore = asyncio.Semaphore(args.limit_model_concurrency)
- await model_semaphore.acquire()
- worker.send_heart_beat()
- generator = worker.generate_stream_gate(params)
- background_tasks = BackgroundTasks()
- background_tasks.add_task(partial(release_model_semaphore, fn=worker.send_heart_beat))
- return StreamingResponse(generator, background=background_tasks)
-
-
-@app.post("/worker_get_status")
-async def get_status(request: Request):
- return worker.get_status()
-
-
-if __name__ == "__main__":
- parser = argparse.ArgumentParser()
- parser.add_argument("--host", type=str, default="localhost")
- parser.add_argument("--port", type=int, default=21002)
- parser.add_argument("--worker-address", type=str,
- default="http://localhost:21002")
- parser.add_argument("--controller-address", type=str,
- default="http://localhost:21001")
- parser.add_argument("--model-path", type=str, default="facebook/opt-350m")
- parser.add_argument("--model-base", type=str, default=None)
- parser.add_argument("--model-name", type=str)
- parser.add_argument("--device", type=str, default="cuda")
- parser.add_argument("--multi-modal", action="store_true", help="Multimodal mode is automatically detected with model name, please make sure `llava` is included in the model path.")
- parser.add_argument("--limit-model-concurrency", type=int, default=5)
- parser.add_argument("--stream-interval", type=int, default=1)
- parser.add_argument("--no-register", action="store_true")
- parser.add_argument("--load-8bit", action="store_true")
- parser.add_argument("--load-4bit", action="store_true")
- args = parser.parse_args()
- logger.info(f"args: {args}")
-
- if args.multi_modal:
- logger.warning("Multimodal mode is automatically detected with model name, please make sure `llava` is included in the model path.")
-
- worker = ModelWorker(args.controller_address,
- args.worker_address,
- worker_id,
- args.no_register,
- args.model_path,
- args.model_base,
- args.model_name,
- args.load_8bit,
- args.load_4bit,
- args.device)
- uvicorn.run(app, host=args.host, port=args.port, log_level="info")
diff --git a/LLaVA-Med/llava/serve/register_worker.py b/LLaVA-Med/llava/serve/register_worker.py
deleted file mode 100644
index 2c2c40295e0351f25709ba25554c9329f15bf0d2..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/serve/register_worker.py
+++ /dev/null
@@ -1,26 +0,0 @@
-"""
-Manually register workers.
-
-Usage:
-python3 -m fastchat.serve.register_worker --controller http://localhost:21001 --worker-name http://localhost:21002
-"""
-
-import argparse
-
-import requests
-
-if __name__ == "__main__":
- parser = argparse.ArgumentParser()
- parser.add_argument("--controller-address", type=str)
- parser.add_argument("--worker-name", type=str)
- parser.add_argument("--check-heart-beat", action="store_true")
- args = parser.parse_args()
-
- url = args.controller_address + "/register_worker"
- data = {
- "worker_name": args.worker_name,
- "check_heart_beat": args.check_heart_beat,
- "worker_status": None,
- }
- r = requests.post(url, json=data)
- assert r.status_code == 200
diff --git a/LLaVA-Med/llava/serve/test_message.py b/LLaVA-Med/llava/serve/test_message.py
deleted file mode 100644
index 8a9b1878833a3da56231e8da1bf877dc70b909d2..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/serve/test_message.py
+++ /dev/null
@@ -1,62 +0,0 @@
-import argparse
-import json
-
-import requests
-
-from llava.conversation import conv_templates
-
-
-def main():
- if args.worker_address:
- worker_addr = args.worker_address
- else:
- controller_addr = args.controller_address
- ret = requests.post(controller_addr + "/refresh_all_workers")
- ret = requests.post(controller_addr + "/list_models")
- models = ret.json()["models"]
- models.sort()
- print(f"Models: {models}")
-
- ret = requests.post(controller_addr + "/get_worker_address",
- json={"model": args.model_name})
- worker_addr = ret.json()["address"]
- print(f"worker_addr: {worker_addr}")
-
- if worker_addr == "":
- return
-
- conv = conv_templates["mistral_instruct"].copy()
- conv.append_message(conv.roles[0], args.message)
- prompt = conv.get_prompt()
-
- headers = {"User-Agent": "LLaVA Client"}
- pload = {
- "model": args.model_name,
- "prompt": prompt,
- "max_new_tokens": args.max_new_tokens,
- "temperature": 0.7,
- "stop": conv.sep2,
- }
- response = requests.post(worker_addr + "/worker_generate_stream", headers=headers,
- json=pload, stream=True)
-
- print(prompt, end="")
- for chunk in response.iter_lines(chunk_size=8192, decode_unicode=False, delimiter=b"\0"):
- if chunk:
- data = json.loads(chunk.decode("utf-8"))
- output = data["text"].split("[/INST]")[-1]
- print(output, end="\r")
- print("")
-
-
-if __name__ == "__main__":
- parser = argparse.ArgumentParser()
- parser.add_argument("--controller-address", type=str, default="http://localhost:21001")
- parser.add_argument("--worker-address", type=str)
- parser.add_argument("--model-name", type=str, default="facebook/opt-350m")
- parser.add_argument("--max-new-tokens", type=int, default=256)
- parser.add_argument("--message", type=str, default=
- "Tell me a story with more than 1000 words.")
- args = parser.parse_args()
-
- main()
diff --git a/LLaVA-Med/llava/utils.py b/LLaVA-Med/llava/utils.py
deleted file mode 100644
index 4006cf917e26c365080b0844c56fab78c48457c0..0000000000000000000000000000000000000000
--- a/LLaVA-Med/llava/utils.py
+++ /dev/null
@@ -1,126 +0,0 @@
-import datetime
-import logging
-import logging.handlers
-import os
-import sys
-
-import requests
-
-from llava.constants import LOGDIR
-
-server_error_msg = "**NETWORK ERROR DUE TO HIGH TRAFFIC. PLEASE REGENERATE OR REFRESH THIS PAGE.**"
-moderation_msg = "YOUR INPUT VIOLATES OUR CONTENT MODERATION GUIDELINES. PLEASE TRY AGAIN."
-
-handler = None
-
-
-def build_logger(logger_name, logger_filename):
- global handler
-
- formatter = logging.Formatter(
- fmt="%(asctime)s | %(levelname)s | %(name)s | %(message)s",
- datefmt="%Y-%m-%d %H:%M:%S",
- )
-
- # Set the format of root handlers
- if not logging.getLogger().handlers:
- logging.basicConfig(level=logging.INFO)
- logging.getLogger().handlers[0].setFormatter(formatter)
-
- # Redirect stdout and stderr to loggers
- stdout_logger = logging.getLogger("stdout")
- stdout_logger.setLevel(logging.INFO)
- sl = StreamToLogger(stdout_logger, logging.INFO)
- sys.stdout = sl
-
- stderr_logger = logging.getLogger("stderr")
- stderr_logger.setLevel(logging.ERROR)
- sl = StreamToLogger(stderr_logger, logging.ERROR)
- sys.stderr = sl
-
- # Get logger
- logger = logging.getLogger(logger_name)
- logger.setLevel(logging.INFO)
-
- # Add a file handler for all loggers
- if handler is None:
- os.makedirs(LOGDIR, exist_ok=True)
- filename = os.path.join(LOGDIR, logger_filename)
- handler = logging.handlers.TimedRotatingFileHandler(
- filename, when='D', utc=True, encoding='UTF-8')
- handler.setFormatter(formatter)
-
- for name, item in logging.root.manager.loggerDict.items():
- if isinstance(item, logging.Logger):
- item.addHandler(handler)
-
- return logger
-
-
-class StreamToLogger(object):
- """
- Fake file-like stream object that redirects writes to a logger instance.
- """
- def __init__(self, logger, log_level=logging.INFO):
- self.terminal = sys.stdout
- self.logger = logger
- self.log_level = log_level
- self.linebuf = ''
-
- def __getattr__(self, attr):
- return getattr(self.terminal, attr)
-
- def write(self, buf):
- temp_linebuf = self.linebuf + buf
- self.linebuf = ''
- for line in temp_linebuf.splitlines(True):
- # From the io.TextIOWrapper docs:
- # On output, if newline is None, any '\n' characters written
- # are translated to the system default line separator.
- # By default sys.stdout.write() expects '\n' newlines and then
- # translates them so this is still cross platform.
- if line[-1] == '\n':
- self.logger.log(self.log_level, line.rstrip())
- else:
- self.linebuf += line
-
- def flush(self):
- if self.linebuf != '':
- self.logger.log(self.log_level, self.linebuf.rstrip())
- self.linebuf = ''
-
-
-def disable_torch_init():
- """
- Disable the redundant torch default initialization to accelerate model creation.
- """
- import torch
- setattr(torch.nn.Linear, "reset_parameters", lambda self: None)
- setattr(torch.nn.LayerNorm, "reset_parameters", lambda self: None)
-
-
-def violates_moderation(text):
- """
- Check whether the text violates OpenAI moderation API.
- """
- url = "https://api.openai.com/v1/moderations"
- headers = {"Content-Type": "application/json",
- "Authorization": "Bearer " + os.environ["OPENAI_API_KEY"]}
- text = text.replace("\n", "")
- data = "{" + '"input": ' + f'"{text}"' + "}"
- data = data.encode("utf-8")
- try:
- ret = requests.post(url, headers=headers, data=data, timeout=5)
- flagged = ret.json()["results"][0]["flagged"]
- except requests.exceptions.RequestException as e:
- flagged = False
- except KeyError as e:
- flagged = False
-
- return flagged
-
-
-def pretty_print_semaphore(semaphore):
- if semaphore is None:
- return "None"
- return f"Semaphore(value={semaphore._value}, locked={semaphore.locked()})"
diff --git a/LLaVA-Med/pyproject.toml b/LLaVA-Med/pyproject.toml
deleted file mode 100644
index 98bb518e440a16150a695561fb0ce16bd0dc2941..0000000000000000000000000000000000000000
--- a/LLaVA-Med/pyproject.toml
+++ /dev/null
@@ -1,98 +0,0 @@
-[build-system]
-requires = ["setuptools>=61.0"]
-build-backend = "setuptools.build_meta"
-
-[project]
-name = "llava_med"
-version = "1.5.0"
-description = "Towards GPT-4 like large language and visual assistant."
-readme = "README.md"
-requires-python = ">=3.8"
-classifiers = [
- "Programming Language :: Python :: 3",
- "License :: OSI Approved :: Apache Software License",
-]
-dependencies = [
- "transformers==4.36.2",
- "tokenizers>=0.15.0",
- "sentencepiece==0.1.99",
- "shortuuid",
- "accelerate==0.21.0",
- "peft==0.4.0",
- "bitsandbytes==0.41.0",
- "pydantic<2,>=1",
- "markdown2[all]",
- "protobuf",
- "numpy",
- "scikit-learn==1.2.2",
- "gradio==3.35.2",
- "gradio_client==0.2.9",
- "requests",
- "httpx==0.24.0",
- "uvicorn",
- "fastapi",
- "einops==0.6.1",
- "einops-exts==0.0.4",
- "timm==0.9.12",
- "tiktoken",
- "openai==1.12.0",
- "backoff",
-]
-
-[project.optional-dependencies]
-train = ["deepspeed==0.9.5", "ninja", "wandb"]
-eval = [
- "azure-ai-ml",
- "datasets",
- "fire",
- "opencv-python",
- "openpyxl==3.1.2",
- "pillow==9.4.0",
- "python-Levenshtein",
- "rich",
- "streamlit==1.29.0",
- "typer[all]",
- "word2number",
-]
-
-[project.urls]
-"Homepage" = "https://github.com/microsoft/LLaVA-Med"
-"Bug Tracker" = "https://github.com/microsoft/LLaVA-Med/issues"
-
-[tool.setuptools.packages.find]
-exclude = [
- "assets*",
- "benchmark*",
- "docs",
- "dist*",
- "playground*",
- "scripts*",
- "tests*",
-]
-
-[tool.wheel]
-exclude = [
- "assets*",
- "benchmark*",
- "docs",
- "dist*",
- "playground*",
- "scripts*",
- "tests*",
-]
-
-[tool.black]
-line-length = 120
-skip-string-normalization = true
-
-[tool.pyright]
-exclude = [
- "**/__pycache__",
- "playground",
- "_results",
- "_data",
- "models",
- "checkpoints",
- "wandb",
- "docs",
-]







